DNA Replication
- Reference work entry
- Cite this reference work entry

- Zoi Lygerou 5 ,
- K. K. Koutroumpas 6 &
- John Lygeros 6
283 Accesses
DNA synthesis
DNA replication is the process of making an identical copy of the genetic material within each cell (Alberts et al. 2007 ; DePamphilis 2006 ). In eukaryotes, DNA replication takes place during a defined period of the cell cycle , called S (for synthesis) phase. DNA replication must be carried out with great precision every time the cell divides, so that genetic information is preserved. Control mechanisms ensure that every base of the genome is replicated once and only once per cell cycle, thereby safeguarding genomic integrity.

Characteristics
We present key characteristics of DNA replication in eukaryotic cells.
Replication Forks Move Continuously Along the Genome as Replisomes Catalyze DNA Synthesis
Each cell must accurately copy millions of bases of DNA (six billion base pairs in a human cell) before every cell division. DNA replication initiates from thousands of sites along eukaryotic chromosomes, called replication origins . Unwinding of the...
This is a preview of subscription content, log in via an institution to check access.
Access this chapter
- Available as PDF
- Read on any device
- Instant download
- Own it forever
- Available as EPUB and PDF
- Durable hardcover edition
- Dispatched in 3 to 5 business days
- Free shipping worldwide - see info
Tax calculation will be finalised at checkout
Purchases are for personal use only
Institutional subscriptions
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2007) Molecular biology of the cell, 5th edn. Garland Science, New York
Google Scholar
Blow JJ, Dutta A (2005) Preventing re-replication of chromosomal DNA. Nat Rev Mol Cell Biol 6:476–486
Article PubMed CAS Google Scholar
Blow JJ, Ge XQ (2009) A model for DNA replication showing how dormant origins safeguard against replication fork failure. EMBO Rep 10:406–412
de Moura APS, Retkute R, Hawkins M, Nieduszynski CA (2010) Mathematical modelling of whole chromosome replication. Nucleic Acids Res 38:5623–5633
DePamphilis ML (2006) DNA replication and human disease. CSHL Press, Cold Spring Harbor
Gilbert DM (2004) In search of the holy replicator. Nat Rev Mol Cell Biol 5:848–855
Herrick J, Jun S, Bechhoefer J, Bensimon A (2002) Kinetic model of DNA replication in eukaryotic organisms. J Mol Biol 320:741–750
Hyrien O, Goldar A (2010) Mathematical modelling of eukaryotic DNA replication. Chromosome Res 18:147–161
Legouras I, Xouri G, Dimopoulos S, Lygeros J, Lygerou Z (2006) DNA replication in the fission yeast: robustness in the face of uncertainty. Yeast 23:951–962
Lygeros J, Koutroumpas K, Dimopoulos S, Legouras I, Kouretas P, Heichinger C, Nurse P, Lygerou Z (2008) Stochastic hybrid modeling of DNA replication across a complete genome. Proc Natl Acad Sci USA 105:12295–300
Spiesser TW, Klipp E, Barberis M (2009) A model for the spatiotemporal organization of DNA replication in Saccharomyces cerevisiae . Mol Genet Genomics 282:25–35
Yang SC, Rhind N, Bechhoefer J (2010) Modeling genome-wide replication kinetics reveals a mechanism for regulation of replication timing. Mol Syst Biol 6:404
PubMed Google Scholar
Download references
Author information
Authors and affiliations.
School of Medicine, Laboratory of General Biology, University of Patras, Rio, Patras, 26500, Greece
Dr. Zoi Lygerou
Automatic Control Laboratory, ETH Zurich, ETL I 22, Physikstrasse 3, Zurich, 8092, Switzerland
K. K. Koutroumpas & Dr. John Lygeros
You can also search for this author in PubMed Google Scholar
Corresponding authors
Correspondence to Zoi Lygerou or John Lygeros .
Editor information
Editors and affiliations.
Biomedical Sciences Research Institute, University of Ulster, Coleraine, UK
Werner Dubitzky
Department of Computer Science, University of Rostock, Rostock, Germany
Olaf Wolkenhauer
Department of Bio and Brain Engineering, Korea Advanced Institute of Science and Technology (KAIST), Daejeon, Republic of Korea
Kwang-Hyun Cho
Department of Biomedical Engineering, Rensselaer Polytechnic Institute, Troy, NY, USA
Hiroki Yokota
Rights and permissions
Reprints and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this entry
Cite this entry.
Lygerou, Z., Koutroumpas, K.K., Lygeros, J. (2013). DNA Replication. In: Dubitzky, W., Wolkenhauer, O., Cho, KH., Yokota, H. (eds) Encyclopedia of Systems Biology. Springer, New York, NY. https://doi.org/10.1007/978-1-4419-9863-7_40
Download citation
DOI : https://doi.org/10.1007/978-1-4419-9863-7_40
Publisher Name : Springer, New York, NY
Print ISBN : 978-1-4419-9862-0
Online ISBN : 978-1-4419-9863-7
eBook Packages : Biomedical and Life Sciences Reference Module Biomedical and Life Sciences
Share this entry
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
DNA replication articles within Nature Reviews Genetics
Research Highlight | 27 September 2022
Promoting a new view of mitochondrial genome regulation
A paper in Molecular Cell reports the characterization of a second functional light strand promoter (LSP2) in the mitochondrial genome, challenging the view that mitochondrial DNA replication and gene expression are coupled by their reliance on a single light strand promoter (LSP).
- Dorothy Clyde
In Brief | 17 March 2022
DNA replication in cell fate reprogramming
A recent study in Nature Genetics investigates the role of DNA replication in cellular plasticity during mouse embryonic development.
Review Article | 23 June 2021
The origin of human mutation in light of genomic data
Genome-scale sequencing data have revealed statistical properties of mutagenesis in humans. Statistical analyses that interpret these patterns and incorporate knowledge on DNA replication and repair pathways can provide mechanistic models that shed light on the origin of spontaneous human mutation in the germ line.
- Vladimir B. Seplyarskiy
- & Shamil Sunyaev
Review Article | 25 November 2019
Chromosome organization in bacteria: mechanistic insights into genome structure and function
Advances in sequencing- and imaging-based techniques for chromosome structure analysis have led to a mature understanding of bacterial chromosome structure and dynamics. In this Review, Dame, Rashid and Grainger discuss the hierarchical nature of bacterial chromosome structure and how it is influenced by diverse types of nucleoid-associated proteins. Furthermore, they describe roles for nucleoid-associated proteins and chromosome structure, including in gene expression, chromosome segregation and cell cycle regulation.
- Remus T. Dame
- , Fatema-Zahra M. Rashid
- & David C. Grainger
Review Article | 08 May 2019
Molecular digital data storage using DNA
Throughout evolution, DNA has been the primary medium of biological information storage. In this article, Ceze, Nivala and Strauss discuss how DNA can be adopted as a storage medium for custom data, as a potential future complement to current data storage media such as computer hard disks, optical disks and tape. They discuss strategies for coding, decoding and error correction and give examples of implementation both in vitro and in vivo.
- , Jeff Nivala
- & Karin Strauss
Perspective | 13 February 2019
Telomeres and telomerase: three decades of progress
In this Timeline article, Shay and Wright provide a historical account of progress in our understanding of telomeres (the ends of linear chromosomes) and telomerase (the primary enzyme that maintains and extends telomere lengths). Their perspective covers seminal moments from the early discoveries through to our latest understanding of the roles of telomeres and telomerase in ageing, diverse human diseases and gene regulation.
- Jerry W. Shay
- & Woodring E. Wright
Research Highlight | 27 November 2017
Multiplex genome engineering in eukaryotes
- Katharine H. Wrighton
Review Article | 17 July 2017
The impact of replication stress on replication dynamics and DNA damage in vertebrate cells
Recent studies have provided insights into the sources of endogenous replication stress, which can result in DNA damage, checkpoint activation and genome-wide replication fork slowing. The authors review established mechanisms involved in the replication stress response, and propose a new model that reconciles data gained from different cellular models.
- Hervé Técher
- , Stéphane Koundrioukoff
- & Michelle Debatisse
Review Article | 21 November 2016
Order from clutter: selective interactions at mammalian replication origins
Genome-wide mapping, mathematical models and functional genetic analyses suggest that distinct molecular interactions at replication initiation sites underlie the regulation of DNA replication in metazoans. In this Review, the authors discuss recent insights into these DNA–protein interactions, and the genetic and epigenetic features of mammalian replication origins.
- Mirit I. Aladjem
- & Christophe E. Redon
Review Article | 29 February 2016
Mechanisms underlying structural variant formation in genomic disorders
Recent studies have revealed a ubiquitous role for genome architecture in the formation of structural variants at a given locus, both in DNA recombination-based and in DNA replication-based processes. These reports showcase the influence of repeat sequences on genomic stability and structural variant complexity and the tremendous plasticity and dynamic nature of our genome.
- Claudia M. B. Carvalho
- & James R. Lupski
Research Highlight | 05 January 2016
The cell cycle flavours of repair
- Eytan Zlotorynski
Research Highlight | 14 December 2015
Quad-jumping
Review Article | 15 September 2015
R loops: new modulators of genome dynamics and function
R loops form when a transcript hybridizes to a complementary DNA locus to result in an RNA–DNA hybrid and a displaced single DNA strand. Such structures can have detrimental cellular roles by causing genome instability. However, recent studies have provided detailed views of genome-wide R-loop occurrences and uncovered various apparently beneficial roles in gene regulation. This Review discusses our latest understanding of the contrasting functions of R loops and the implications for genome regulation and various diseases.
- José M. Santos-Pereira
- & Andrés Aguilera
Research Highlight | 18 June 2015
Chromothripsis and micronucleus formation
- Denise Waldron
Review Article | 01 July 2014
Mechanisms underlying mutational signatures in human cancers
Mutagenic processes leave characteristic imprints on the cancer genome that can help to identify the underlying DNA damaging components as well as DNA repair and replicative pathways that are active or disrupted. This Review discusses these mutational signatures according to different classes of mutations and summarizes how different components contribute mechanistically to produce each signature type.
- Thomas Helleday
- , Saeed Eshtad
- & Serena Nik-Zainal
Research Highlight | 24 April 2014
Amplified origins of antibiotic resistance
- Darren J. Burgess
Research Highlight | 26 November 2013
Organ roles for mitochondrial mutations?
Research Highlight | 13 August 2013
Replication factors make the transition
- Hannah Stower
In Brief | 16 January 2013
ChIP–seq for human replication origins
In Brief | 18 December 2012
A U-turn for mutagenesis?
In Brief | 18 September 2012
Chromatin inheritance during DNA replication
Review Article | 14 February 2012
Transcription as a source of genome instability
Transcription poses a risk to the genome through transcription-associated mutagenesis and recombination. This Review discusses recent findings about influences on this genomic instability, such as the rate and direction of transcription or nucleic acid structures, and how these phenomena may be considered across species.
- & Sue Jinks-Robertson
In Brief | 17 January 2012
Asymmetry caused by replication-coupled chromatin assembly
Review Article | 01 September 2010
Evaluating genome-scale approaches to eukaryotic DNA replication
This article reviews the increasing range of genome-scale methods that are being used to analyse eukaryotic DNA replication. Studies in different species and of replication timing or origin location have yielded varying degrees of success; technical hurdles remain, but important biological insights have been gained.
- David M. Gilbert
Browse broader subjects
- Molecular biology
- Cell division
Browse narrower subjects
- DNA synthesis
- Fragile sites
- Origin firing
- Origin selection
- Stalled forks
- Translesion synthesis
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies

- school Campus Bookshelves
- menu_book Bookshelves
- perm_media Learning Objects
- login Login
- how_to_reg Request Instructor Account
- hub Instructor Commons
- Download Page (PDF)
- Download Full Book (PDF)
- Periodic Table
- Physics Constants
- Scientific Calculator
- Reference & Cite
- Tools expand_more
- Readability
selected template will load here
This action is not available.

11.2: DNA Replication
- Last updated
- Save as PDF
- Page ID 5181

Learning Objectives
- Explain the meaning of semiconservative DNA replication
- Explain why DNA replication is bidirectional and includes both a leading and lagging strand
- Explain why Okazaki fragments are formed
- Describe the process of DNA replication and the functions of the enzymes involved
- Identify the differences between DNA replication in bacteria and eukaryotes
- Explain the process of rolling circle replication
The elucidation of the structure of the double helix by James Watson and Francis Crick in 1953 provided a hint as to how DNA is copied during the process of replication. Separating the strands of the double helix would provide two templates for the synthesis of new complementary strands, but exactly how new DNA molecules were constructed was still unclear. In one model, semiconservative replication, the two strands of the double helix separate during DNA replication, and each strand serves as a template from which the new complementary strand is copied; after replication, each double-stranded DNA includes one parental or “old” strand and one “new” strand. There were two competing models also suggested: conservative and dispersive, which are shown in Figure \(\PageIndex{1}\).
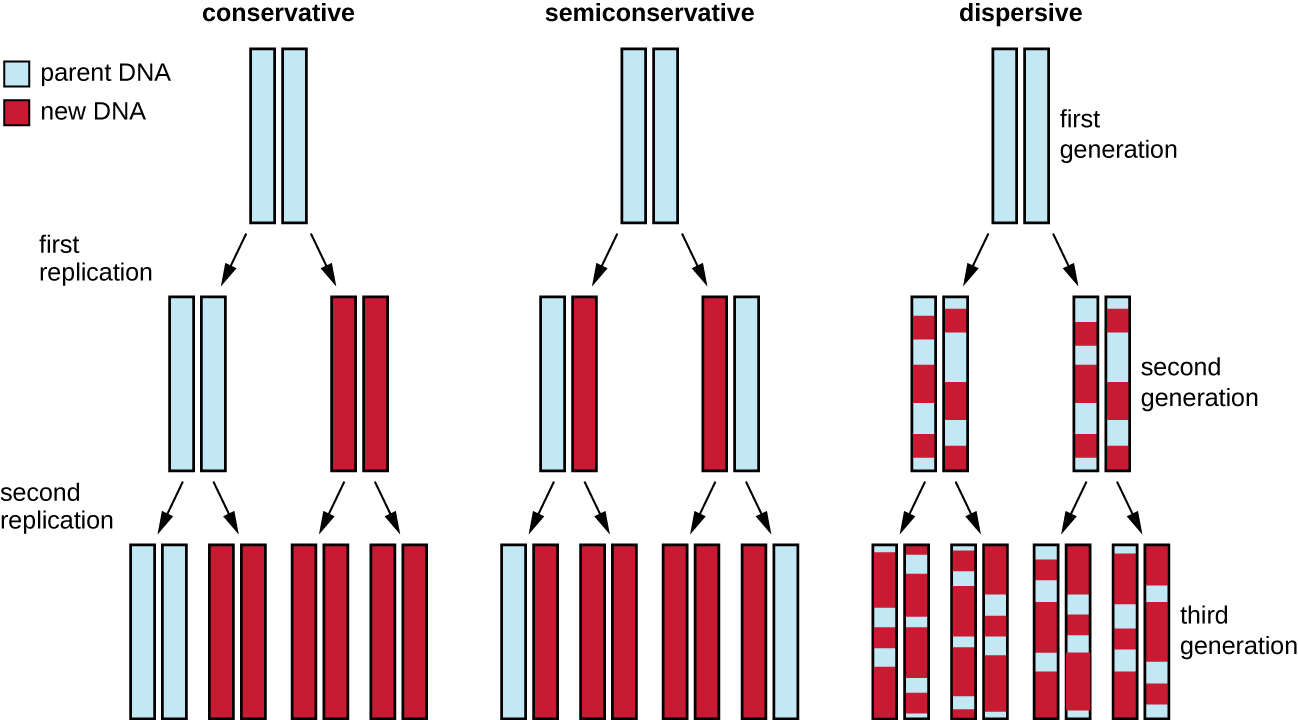
Matthew Meselson (1930–) and Franklin Stahl (1929–) devised an experiment in 1958 to test which of these models correctly represents DNA replication (Figure \(\PageIndex{2}\)). They grew E. coli for several generations in a medium containing a “heavy” isotope of nitrogen ( 15 N) that was incorporated into nitrogenous bases and, eventually, into the DNA. This labeled the parental DNA. The E. coli culture was then shifted into a medium containing 14 N and allowed to grow for one generation. The cells were harvested and the DNA was isolated. The DNA was separated by ultracentrifugation, during which the DNA formed bands according to its density. DNA grown in 15 N would be expected to form a band at a higher density position than that grown in 14 N. Meselson and Stahl noted that after one generation of growth in 14 N, the single band observed was intermediate in position in between DNA of cells grown exclusively in 15 N or 14 N. This suggested either a semiconservative or dispersive mode of replication. Some cells were allowed to grow for one more generation in 14 N and spun again. The DNA harvested from cells grown for two generations in 14 N formed two bands: one DNA band was at the intermediate position between 15 N and 14 N, and the other corresponded to the band of 14 N DNA. These results could only be explained if DNA replicates in a semiconservative manner. Therefore, the other two models were ruled out. As a result of this experiment, we now know that during DNA replication, each of the two strands that make up the double helix serves as a template from which new strands are copied. The new strand will be complementary to the parental or “old” strand. The resulting DNA molecules have the same sequence and are divided equally into the two daughter cells.
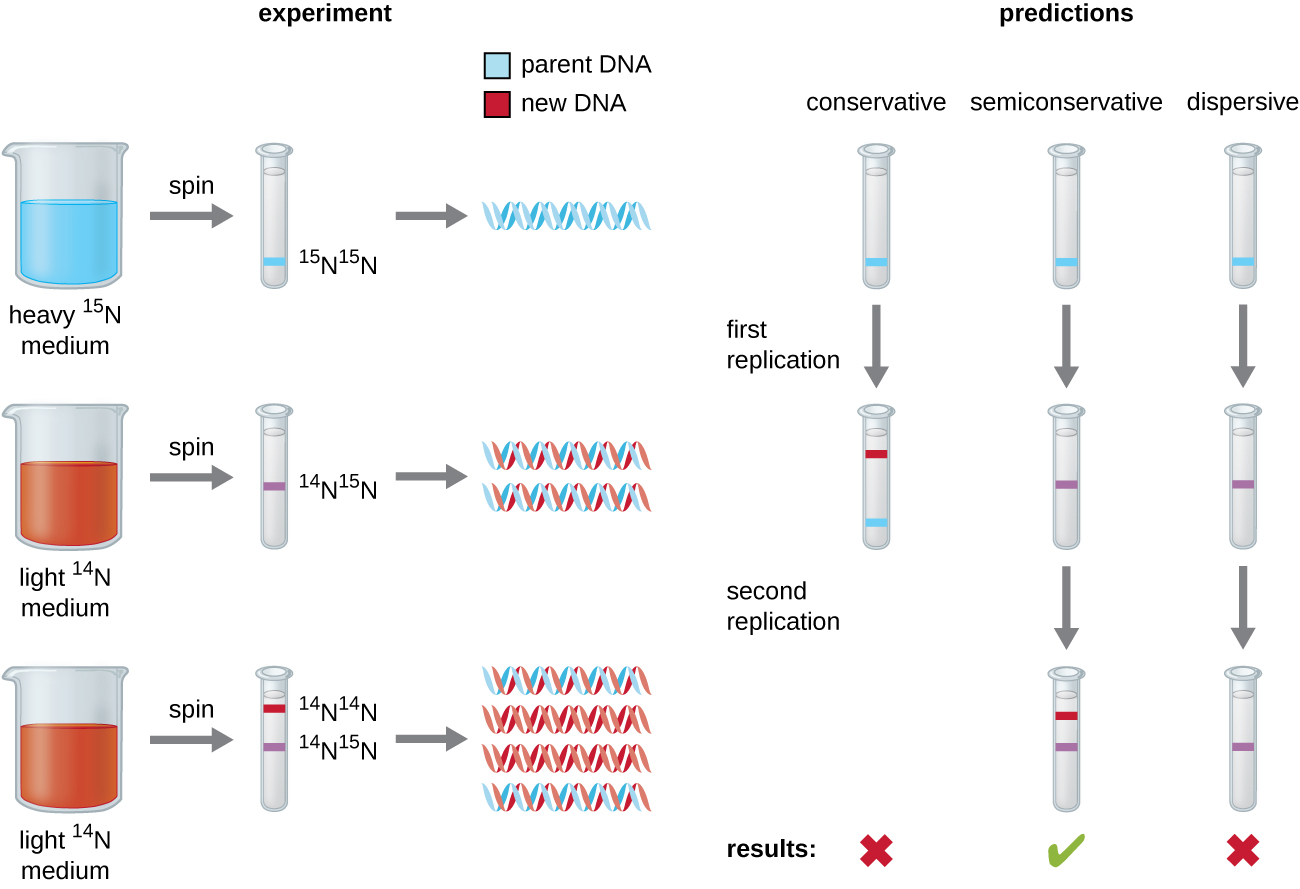

Exercise \(\PageIndex{1}\)
What would have been the conclusion of Meselson and Stahl’s experiment if, after the first generation, they had found two bands of DNA?
DNA Replication in Bacteria
DNA replication has been well studied in bacteria primarily because of the small size of the genome and the mutants that are available. E. coli has 4.6 million base pairs (Mbp) in a single circular chromosome and all of it is replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the circle bidirectionally (i.e., in both directions). This means that approximately 1000 nucleotides are added per second. The process is quite rapid and occurs with few errors.
DNA replication uses a large number of proteins and enzymes (Table \(\PageIndex{1}\)). One of the key players is the enzyme DNA polymerase, also known as DNA pol. In bacteria, three main types of DNA polymerases are known: DNA pol I, DNA pol II, and DNA pol III. It is now known that DNA pol III is the enzyme required for DNA synthesis; DNA pol I and DNA pol II are primarily required for repair. DNA pol III adds deoxyribonucleotides each complementary to a nucleotide on the template strand, one by one to the 3’-OH group of the growing DNA chain. The addition of these nucleotides requires energy. This energy is present in the bonds of three phosphate groups attached to each nucleotide (a triphosphate nucleotide), similar to how energy is stored in the phosphate bonds of adenosine triphosphate (ATP) (Figure \(\PageIndex{3}\)). When the bond between the phosphates is broken and diphosphate is released, the energy released allows for the formation of a covalent phosphodiester bond by dehydration synthesis between the incoming nucleotide and the free 3’-OH group on the growing DNA strand.
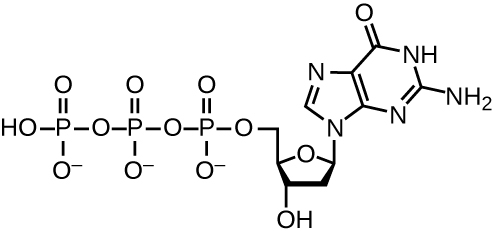
The initiation of replication occurs at specific nucleotide sequence called the origin of replication, where various proteins bind to begin the replication process. E. coli has a single origin of replication (as do most prokaryotes), called oriC , on its one chromosome. The origin of replication is approximately 245 base pairs long and is rich in adenine-thymine (AT) sequences.
Some of the proteins that bind to the origin of replication are important in making single-stranded regions of DNA accessible for replication. Chromosomal DNA is typically wrapped around histones (in eukaryotes and archaea) or histone-like proteins (in bacteria), and is supercoiled, or extensively wrapped and twisted on itself. This packaging makes the information in the DNA molecule inaccessible. However, enzymes called topoisomerases change the shape and supercoiling of the chromosome. For bacterial DNA replication to begin, the supercoiled chromosome is relaxed by topoisomerase II, also called DNA gyrase. An enzyme called helicase then separates the DNA strands by breaking the hydrogen bonds between the nitrogenous base pairs. Recall that AT sequences have fewer hydrogen bonds and, hence, have weaker interactions than guanine-cytosine (GC) sequences. These enzymes require ATP hydrolysis. As the DNA opens up, Y-shaped structures called replication forks are formed. Two replication forks are formed at the origin of replication, allowing for bidirectional replication and formation of a structure that looks like a bubble when viewed with a transmission electron microscope; as a result, this structure is called a replication bubble. The DNA near each replication fork is coated with single-stranded binding proteins to prevent the single-stranded DNA from rewinding into a double helix.
Once single-stranded DNA is accessible at the origin of replication, DNA replication can begin. However, DNA pol III is able to add nucleotides only in the 5’ to 3’ direction (a new DNA strand can be only extended in this direction). This is because DNA polymerase requires a free 3’-OH group to which it can add nucleotides by forming a covalent phosphodiester bond between the 3’-OH end and the 5’ phosphate of the next nucleotide. This also means that it cannot add nucleotides if a free 3’-OH group is not available, which is the case for a single strand of DNA. The problem is solved with the help of an RNA sequence that provides the free 3’-OH end. Because this sequence allows the start of DNA synthesis, it is appropriately called the primer. The primer is five to 10 nucleotides long and complementary to the parental or template DNA. It is synthesized by RNA primase, which is an RNA polymerase. Unlike DNA polymerases, RNA polymerases do not need a free 3’-OH group to synthesize an RNA molecule. Now that the primer provides the free 3’-OH group, DNA polymerase III can now extend this RNA primer, adding DNA nucleotides one by one that are complementary to the template strand (Figure \(\PageIndex{1}\)).
During elongation in DNA replication, the addition of nucleotides occurs at its maximal rate of about 1000 nucleotides per second. DNA polymerase III can only extend in the 5’ to 3’ direction, which poses a problem at the replication fork. The DNA double helix is antiparallel; that is, one strand is oriented in the 5’ to 3’ direction and the other is oriented in the 3’ to 5’ direction (see Structure and Function of DNA ). During replication, one strand, which is complementary to the 3’ to 5’ parental DNA strand, is synthesized continuously toward the replication fork because polymerase can add nucleotides in this direction. This continuously synthesized strand is known as the leading strand. The other strand, complementary to the 5’ to 3’ parental DNA, grows away from the replication fork, so the polymerase must move back toward the replication fork to begin adding bases to a new primer, again in the direction away from the replication fork. It does so until it bumps into the previously synthesized strand and then it moves back again (Figure \(\PageIndex{4}\)). These steps produce small DNA sequence fragments known as Okazaki fragments, each separated by RNA primer. Okazaki fragments are named after the Japanese research team and married couple Reiji and Tsuneko Okazaki, who first discovered them in 1966. The strand with the Okazaki fragments is known as the lagging strand, and its synthesis is said to be discontinuous.
The leading strand can be extended from one primer alone, whereas the lagging strand needs a new primer for each of the short Okazaki fragments. The overall direction of the lagging strand will be 3’ to 5’, and that of the leading strand 5’ to 3’. A protein called the sliding clamp holds the DNA polymerase in place as it continues to add nucleotides. The sliding clamp is a ring-shaped protein that binds to the DNA and holds the polymerase in place. Beyond its role in initiation, topoisomerase also prevents the overwinding of the DNA double helix ahead of the replication fork as the DNA is opening up; it does so by causing temporary nicks in the DNA helix and then resealing it. As synthesis proceeds, the RNA primers are replaced by DNA. The primers are removed by the exonuclease activity of DNA polymerase I, and the gaps are filled in. The nicks that remain between the newly synthesized DNA (that replaced the RNA primer) and the previously synthesized DNA are sealed by the enzyme DNA ligase that catalyzes the formation of covalent phosphodiester linkage between the 3’-OH end of one DNA fragment and the 5’ phosphate end of the other fragment, stabilizing the sugar-phosphate backbone of the DNA molecule.
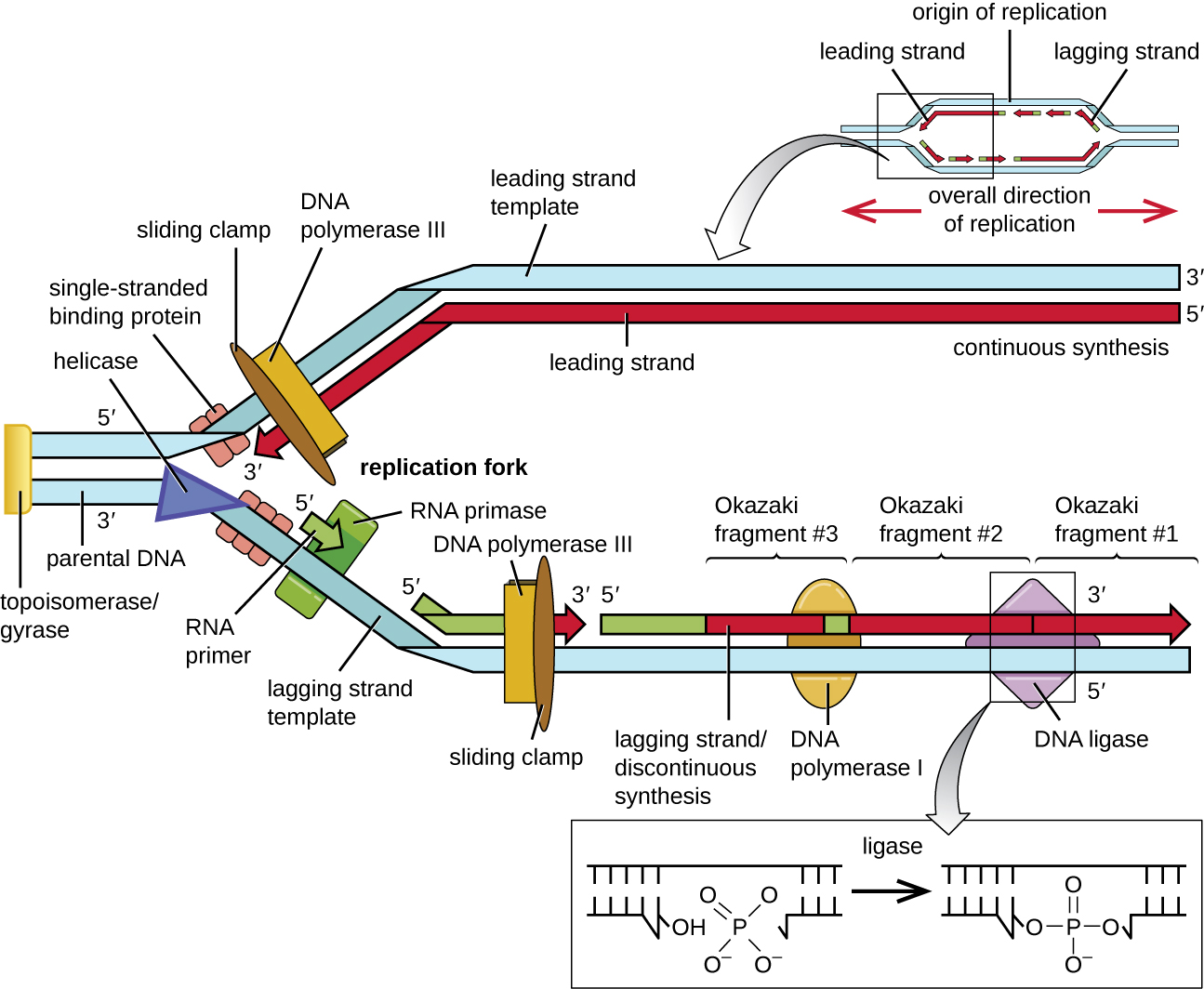
Termination
Once the complete chromosome has been replicated, termination of DNA replication must occur. Although much is known about initiation of replication, less is known about the termination process. Following replication, the resulting complete circular genomes of prokaryotes are concatenated, meaning that the circular DNA chromosomes are interlocked and must be separated from each other. This is accomplished through the activity of bacterial topoisomerase IV, which introduces double-stranded breaks into DNA molecules, allowing them to separate from each other; the enzyme then reseals the circular chromosomes. The resolution of concatemers is an issue unique to prokaryotic DNA replication because of their circular chromosomes. Because both bacterial DNA gyrase and topoisomerase IV are distinct from their eukaryotic counterparts, these enzymes serve as targets for a class of antimicrobial drugs called quinolones.
Exercise \(\PageIndex{2}\)
- Which enzyme breaks the hydrogen bonds holding the two strands of DNA together so that replication can occur?
- Is it the lagging strand or the leading strand that is synthesized in the direction toward the opening of the replication fork?
- Which enzyme is responsible for removing the RNA primers in newly replicated bacterial DNA?
DNA Replication in Eukaryotes
Eukaryotic genomes are much more complex and larger than prokaryotic genomes and are typically composed of multiple linear chromosomes (Table \(\PageIndex{2}\)). The human genome, for example, has 3 billion base pairs per haploid set of chromosomes, and 6 billion base pairs are inserted during replication. There are multiple origins of replication on each eukaryotic chromosome (Figure \(\PageIndex{5}\)); the human genome has 30,000 to 50,000 origins of replication. The rate of replication is approximately 100 nucleotides per second—10 times slower than prokaryotic replication.
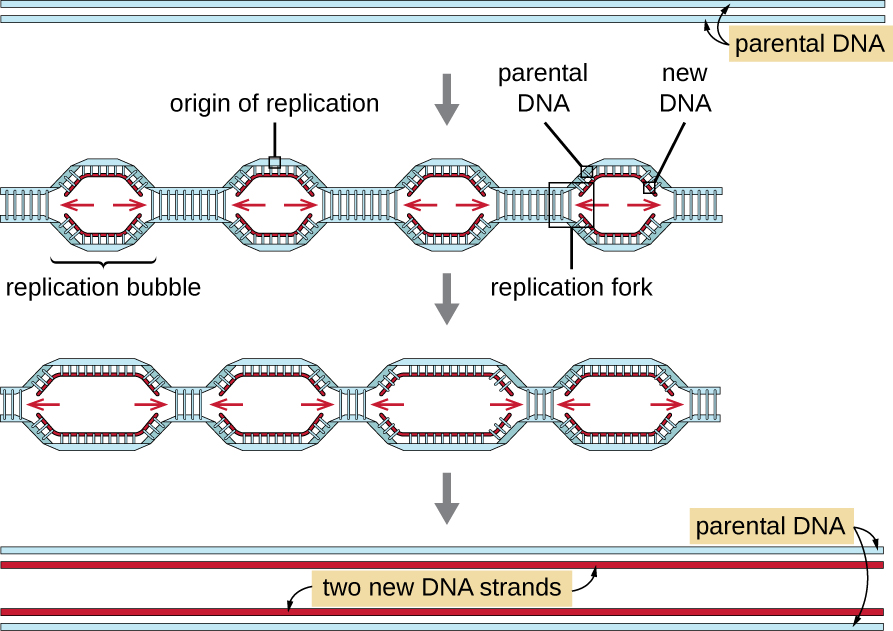
The essential steps of replication in eukaryotes are the same as in prokaryotes. Before replication can start, the DNA has to be made available as a template. Eukaryotic DNA is highly supercoiled and packaged, which is facilitated by many proteins, including histones (see Structure and Function of Cellular Genomes ). At the origin of replication, a prereplication complex composed of several proteins, including helicase, forms and recruits other enzymes involved in the initiation of replication, including topoisomerase to relax supercoiling, single-stranded binding protein, RNA primase, and DNA polymerase. Following initiation of replication, in a process similar to that found in prokaryotes, elongation is facilitated by eukaryotic DNA polymerases. The leading strand is continuously synthesized by the eukaryotic polymerase enzyme pol δ, while the lagging strand is synthesized by pol ε. A sliding clamp protein holds the DNA polymerase in place so that it does not fall off the DNA. The enzyme ribonuclease H (RNase H), instead of a DNA polymerase as in bacteria, removes the RNA primer, which is then replaced with DNA nucleotides. The gaps that remain are sealed by DNA ligase.
Because eukaryotic chromosomes are linear, one might expect that their replication would be more straightforward. As in prokaryotes, the eukaryotic DNA polymerase can add nucleotides only in the 5’ to 3’ direction. In the leading strand, synthesis continues until it reaches either the end of the chromosome or another replication fork progressing in the opposite direction. On the lagging strand, DNA is synthesized in short stretches, each of which is initiated by a separate primer. When the replication fork reaches the end of the linear chromosome, there is no place to make a primer for the DNA fragment to be copied at the end of the chromosome. These ends thus remain unpaired and, over time, they may get progressively shorter as cells continue to divide.
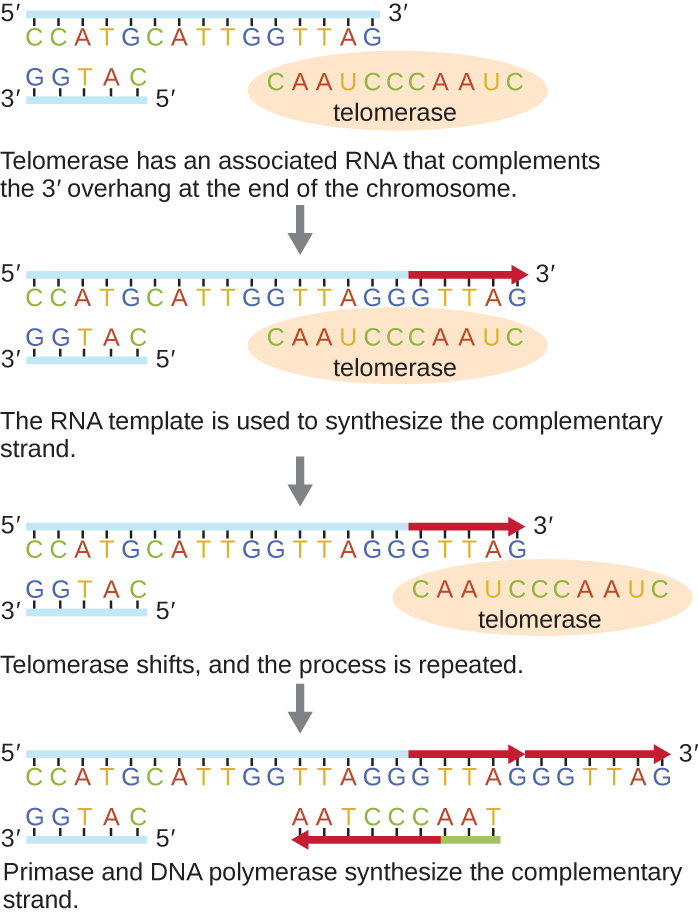
The ends of the linear chromosomes are known as telomeres and consist of noncoding repetitive sequences. The telomeres protect coding sequences from being lost as cells continue to divide. In humans, a six base-pair sequence, TTAGGG, is repeated 100 to 1000 times to form the telomere. The discovery of the enzyme telomerase (Figure \(\PageIndex{6}\)) clarified our understanding of how chromosome ends are maintained. Telomerase contains a catalytic part and a built-in RNA template. It attaches to the end of the chromosome, and complementary bases to the RNA template are added on the 3’ end of the DNA strand. Once the 3’ end of the lagging strand template is sufficiently elongated, DNA polymerase can add the nucleotides complementary to the ends of the chromosomes. In this way, the ends of the chromosomes are replicated. In humans, telomerase is typically active in germ cells and adult stem cells; it is not active in adult somatic cells and may be associated with the aging of these cells. Eukaryotic microbes including fungi and protozoans also produce telomerase to maintain chromosomal integrity. For her discovery of telomerase and its action, Elizabeth Blackburn (1948–) received the Nobel Prize for Medicine or Physiology in 2009.
Exercise \(\PageIndex{3}\)
- How does the origin of replication differ between eukaryotes and prokaryotes?
- What polymerase enzymes are responsible for DNA synthesis during eukaryotic replication?
- What is found at the ends of the chromosomes in eukaryotes and why?
DNA Replication of Extrachromosomal Elements: Plasmids and Viruses
To copy their nucleic acids, plasmids and viruses frequently use variations on the pattern of DNA replication described for prokaryote genomes. For more information on the wide range of viral replication strategies, see The Viral Life Cycle .
Rolling Circle Replication
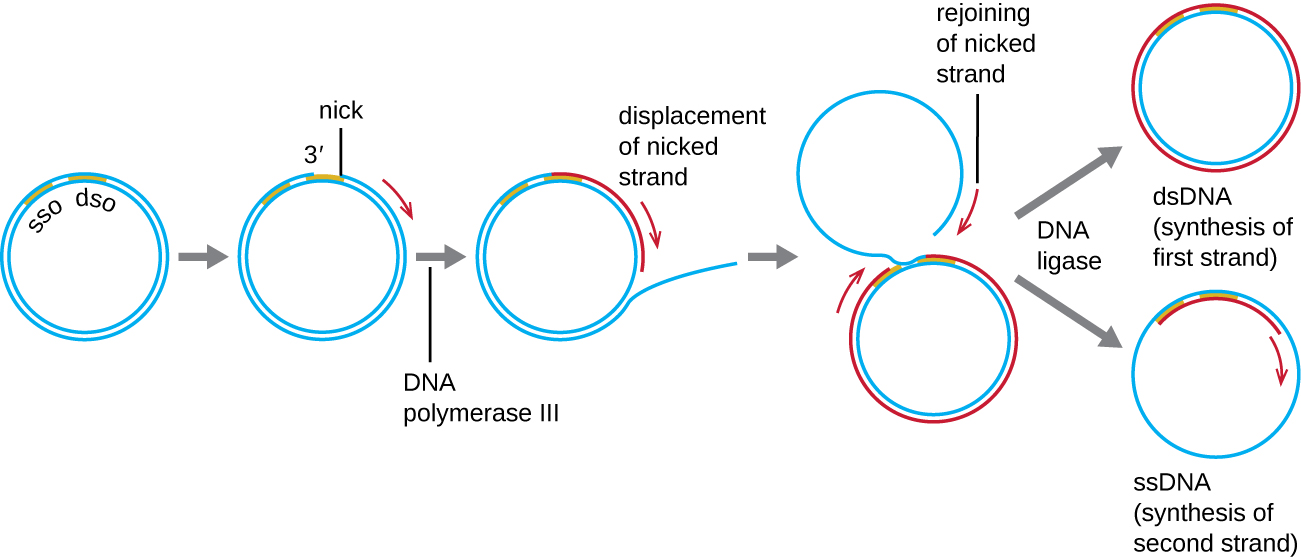
Whereas many bacterial plasmids (see Unique Characteristics of Prokaryotic Cells ) replicate by a process similar to that used to copy the bacterial chromosome, other plasmids, several bacteriophages, and some viruses of eukaryotes use rolling circle replication (Figure \(\PageIndex{7}\)). The circular nature of plasmids and the circularization of some viral genomes on infection make this possible. Rolling circle replication begins with the enzymatic nicking of one strand of the double-stranded circular molecule at the double-stranded origin (dso) site. In bacteria, DNA polymerase III binds to the 3’-OH group of the nicked strand and begins to unidirectionally replicate the DNA using the un-nicked strand as a template, displacing the nicked strand as it does so. Completion of DNA replication at the site of the original nick results in full displacement of the nicked strand, which may then recircularize into a single-stranded DNA molecule. RNA primase then synthesizes a primer to initiate DNA replication at the single-stranded origin (sso) site of the single-stranded DNA (ssDNA) molecule, resulting in a double-stranded DNA (dsDNA) molecule identical to the other circular DNA molecule.
Exercise \(\PageIndex{4}\)
Is there a lagging strand in rolling circle replication? Why or why not?
Key Concepts and Summary
- The DNA replication process is semiconservative , which results in two DNA molecules, each having one parental strand of DNA and one newly synthesized strand.
- In bacteria, the initiation of replication occurs at the origin of replication , where supercoiled DNA is unwound by DNA gyrase , made single-stranded by helicase , and bound by single-stranded binding protein to maintain its single-stranded state. Primase synthesizes a short RNA primer , providing a free 3’-OH group to which DNA polymerase III can add DNA nucleotides.
- During elongation , the leading strand of DNA is synthesized continuously from a single primer. The lagging strand is synthesized discontinuously in short Okazaki fragments , each requiring its own primer. The RNA primers are removed and replaced with DNA nucleotides by bacterial DNA polymerase I , and DNA ligase seals the gaps between these fragments.
- Termination of replication in bacteria involves the resolution of circular DNA concatemers by topoisomerase IV to release the two copies of the circular chromosome.
- Eukaryotes typically have multiple linear chromosomes, each with multiple origins of replication. Overall, replication in eukaryotes is similar to that in prokaryotes.
- The linear nature of eukaryotic chromosomes necessitates telomeres to protect genes near the end of the chromosomes. Telomerase extends telomeres, preventing their degradation, in some cell types.
- Rolling circle replication is a type of rapid unidirectional DNA synthesis of a circular DNA molecule used for the replication of some plasmids.
11.2 DNA Replication
Learning objectives.
By the end of this section, you will be able to:
- Explain the meaning of semiconservative DNA replication
- Explain why DNA replication is bidirectional and includes both a leading and lagging strand
- Explain why Okazaki fragments are formed
- Describe the process of DNA replication and the functions of the enzymes involved
- Identify the differences between DNA replication in bacteria and eukaryotes
- Explain the process of rolling circle replication
The elucidation of the structure of the double helix by James Watson and Francis Crick in 1953 provided a hint as to how DNA is copied during the process of replication . Separating the strands of the double helix would provide two templates for the synthesis of new complementary strands, but exactly how new DNA molecules were constructed was still unclear. In one model, semiconservative replication , the two strands of the double helix separate during DNA replication, and each strand serves as a template from which the new complementary strand is copied; after replication, each double-stranded DNA includes one parental or “old” strand and one “new” strand. There were two competing models also suggested: conservative and dispersive, which are shown in Figure 11.4 .
Matthew Meselson (1930–) and Franklin Stahl (1929–) devised an experiment in 1958 to test which of these models correctly represents DNA replication ( Figure 11.5 ). They grew E. coli for several generations in a medium containing a “heavy” isotope of nitrogen ( 15 N) that was incorporated into nitrogenous bases and, eventually, into the DNA. This labeled the parental DNA. The E. coli culture was then shifted into a medium containing 14 N and allowed to grow for one generation. The cells were harvested and the DNA was isolated. The DNA was separated by ultracentrifugation, during which the DNA formed bands according to its density. DNA grown in 15 N would be expected to form a band at a higher density position than that grown in 14 N. Meselson and Stahl noted that after one generation of growth in 14 N, the single band observed was intermediate in position in between DNA of cells grown exclusively in 15 N or 14 N. This suggested either a semiconservative or dispersive mode of replication. Some cells were allowed to grow for one more generation in 14 N and spun again. The DNA harvested from cells grown for two generations in 14 N formed two bands: one DNA band was at the intermediate position between 15 N and 14 N, and the other corresponded to the band of 14 N DNA. These results could only be explained if DNA replicates in a semiconservative manner. If DNA replication was dispersive, a single purple band positioned closer to the red 14 14 would have been observed, as more 14 was added in a dispersive manner to replace 15 . Therefore, the other two models were ruled out. As a result of this experiment, we now know that during DNA replication, each of the two strands that make up the double helix serves as a template from which new strands are copied. The new strand will be complementary to the parental or “old” strand. The resulting DNA molecules have the same sequence and are divided equally into the two daughter cells.
Check Your Understanding
- What would have been the conclusion of Meselson and Stahl’s experiment if, after the first generation, they had found two bands of DNA?
DNA Replication in Bacteria
DNA replication has been well studied in bacteria primarily because of the small size of the genome and the mutants that are available. E. coli has 4.6 million base pairs (Mbp) in a single circular chromosome and all of it is replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the circle bidirectionally (i.e., in both directions). This means that approximately 1000 nucleotides are added per second. The process is quite rapid and occurs with few errors.
DNA replication uses a large number of proteins and enzymes ( Table 11.1 ). One of the key players is the enzyme DNA polymerase , also known as DNA pol. In bacteria, three main types of DNA polymerases are known: DNA pol I, DNA pol II, and DNA pol III. It is now known that DNA pol III is the enzyme required for DNA synthesis; DNA pol I and DNA pol II are primarily required for repair. DNA pol III adds deoxyribonucleotides each complementary to a nucleotide on the template strand, one by one to the 3’-OH group of the growing DNA chain. The addition of these nucleotides requires energy. This energy is present in the bonds of three phosphate groups attached to each nucleotide (a triphosphate nucleotide), similar to how energy is stored in the phosphate bonds of adenosine triphosphate (ATP) ( Figure 11.6 ). When the bond between the phosphates is broken and diphosphate is released, the energy released allows for the formation of a covalent phosphodiester bond by dehydration synthesis between the incoming nucleotide and the free 3’-OH group on the growing DNA strand.
The initiation of replication occurs at specific nucleotide sequence called the origin of replication , where various proteins bind to begin the replication process. E. coli has a single origin of replication (as do most prokaryotes), called oriC , on its one chromosome. The origin of replication is approximately 245 base pairs long and is rich in adenine-thymine (AT) sequences.
Some of the proteins that bind to the origin of replication are important in making single-stranded regions of DNA accessible for replication. Chromosomal DNA is typically wrapped around histones (in eukaryotes and archaea) or histone-like proteins (in bacteria), and is supercoiled , or extensively wrapped and twisted on itself. This packaging makes the information in the DNA molecule inaccessible. However, enzymes called topoisomerases change the shape and supercoiling of the chromosome. For bacterial DNA replication to begin, the supercoiled chromosome is relaxed by topoisomerase II , also called DNA gyrase . An enzyme called helicase then separates the DNA strands by breaking the hydrogen bonds between the nitrogenous base pairs. Recall that AT sequences have fewer hydrogen bonds and, hence, have weaker interactions than guanine-cytosine (GC) sequences. These enzymes require ATP hydrolysis. As the DNA opens up, Y-shaped structures called replication forks are formed. Two replication forks are formed at the origin of replication, allowing for bidirectional replication and formation of a structure that looks like a bubble when viewed with a transmission electron microscope; as a result, this structure is called a replication bubble . The DNA near each replication fork is coated with single-stranded binding proteins to prevent the single-stranded DNA from rewinding into a double helix.
Once single-stranded DNA is accessible at the origin of replication, DNA replication can begin. However, DNA pol III is able to add nucleotides only in the 5’ to 3’ direction (a new DNA strand can be only extended in this direction). This is because DNA polymerase requires a free 3’-OH group to which it can add nucleotides by forming a covalent phosphodiester bond between the 3’-OH end and the 5’ phosphate of the next nucleotide. This also means that it cannot add nucleotides if a free 3’-OH group is not available, which is the case for a single strand of DNA. The problem is solved with the help of an RNA sequence that provides the free 3’-OH end. Because this sequence allows the start of DNA synthesis, it is appropriately called the primer . The primer is five to 10 nucleotides long and complementary to the parental or template DNA. It is synthesized by RNA primase , which is an RNA polymerase . Unlike DNA polymerases, RNA polymerases do not need a free 3’-OH group to synthesize an RNA molecule. Now that the primer provides the free 3’-OH group, DNA polymerase III can now extend this RNA primer, adding DNA nucleotides one by one that are complementary to the template strand ( Figure 11.4 ).
During elongation in DNA replication , the addition of nucleotides occurs at its maximal rate of about 1000 nucleotides per second. DNA polymerase III can only extend in the 5’ to 3’ direction, which poses a problem at the replication fork. The DNA double helix is antiparallel; that is, one strand is oriented in the 5’ to 3’ direction and the other is oriented in the 3’ to 5’ direction (see Structure and Function of DNA ). During replication, one strand, which is complementary to the 3’ to 5’ parental DNA strand, is synthesized continuously toward the replication fork because polymerase can add nucleotides in this direction. This continuously synthesized strand is known as the leading strand . The other strand, complementary to the 5’ to 3’ parental DNA, grows away from the replication fork, so the polymerase must move back toward the replication fork to begin adding bases to a new primer, again in the direction away from the replication fork. It does so until it bumps into the previously synthesized strand and then it moves back again ( Figure 11.7 ). These steps produce small DNA sequence fragments known as Okazaki fragments , each separated by RNA primer. Okazaki fragments are named after the Japanese research team and married couple Reiji and Tsuneko Okazaki , who first discovered them in 1966. The strand with the Okazaki fragments is known as the lagging strand , and its synthesis is said to be discontinuous.
The leading strand can be extended from one primer alone, whereas the lagging strand needs a new primer for each of the short Okazaki fragments. The overall direction of the lagging strand will be 3’ to 5’, and that of the leading strand 5’ to 3’. A protein called the sliding clamp holds the DNA polymerase in place as it continues to add nucleotides. The sliding clamp is a ring-shaped protein that binds to the DNA and holds the polymerase in place. Beyond its role in initiation, topoisomerase also prevents the overwinding of the DNA double helix ahead of the replication fork as the DNA is opening up; it does so by causing temporary nicks in the DNA helix and then resealing it. As synthesis proceeds, the RNA primers are replaced by DNA. The primers are removed by the exonuclease activity of DNA polymerase I, and the gaps are filled in. The nicks that remain between the newly synthesized DNA (that replaced the RNA primer) and the previously synthesized DNA are sealed by the enzyme DNA ligase that catalyzes the formation of covalent phosphodiester linkage between the 3’-OH end of one DNA fragment and the 5’ phosphate end of the other fragment, stabilizing the sugar-phosphate backbone of the DNA molecule.
Termination
Once the complete chromosome has been replicated, termination of DNA replication must occur. Although much is known about initiation of replication, less is known about the termination process. Following replication, the resulting complete circular genomes of prokaryotes are concatenated, meaning that the circular DNA chromosomes are interlocked and must be separated from each other. This is accomplished through the activity of bacterial topoisomerase IV, which introduces double-stranded breaks into DNA molecules, allowing them to separate from each other; the enzyme then reseals the circular chromosomes. The resolution of concatemers is an issue unique to prokaryotic DNA replication because of their circular chromosomes. Because both bacterial DNA gyrase and topoisomerase IV are distinct from their eukaryotic counterparts, these enzymes serve as targets for a class of antimicrobial drugs called quinolones .
- Which enzyme breaks the hydrogen bonds holding the two strands of DNA together so that replication can occur?
- Is it the lagging strand or the leading strand that is synthesized in the direction toward the opening of the replication fork?
- Which enzyme is responsible for removing the RNA primers in newly replicated bacterial DNA?
DNA Replication in Eukaryotes
Eukaryotic genomes are much more complex and larger than prokaryotic genomes and are typically composed of multiple linear chromosomes ( Table 11.2 ). The human genome , for example, has 3 billion base pairs per haploid set of chromosomes, and 6 billion base pairs are inserted during replication. There are multiple origins of replication on each eukaryotic chromosome ( Figure 11.8 ); the human genome has 30,000 to 50,000 origins of replication. The rate of replication is approximately 100 nucleotides per second—10 times slower than prokaryotic replication.
The essential steps of replication in eukaryotes are the same as in prokaryotes. Before replication can start, the DNA has to be made available as a template. Eukaryotic DNA is highly supercoiled and packaged, which is facilitated by many proteins, including histone s (see Structure and Function of Cellular Genomes ). At the origin of replication , a prereplication complex composed of several proteins, including helicase , forms and recruits other enzymes involved in the initiation of replication, including topoisomerase to relax supercoiling, single-stranded binding protein, RNA primase , and DNA polymerase . Following initiation of replication, in a process similar to that found in prokaryotes, elongation is facilitated by eukaryotic DNA polymerases. The leading strand is continuously synthesized by the eukaryotic polymerase enzyme pol δ, while the lagging strand is synthesized by pol ε. A sliding clamp protein holds the DNA polymerase in place so that it does not fall off the DNA. The enzyme ribonuclease H ( RNase H ), instead of a DNA polymerase as in bacteria, removes the RNA primer, which is then replaced with DNA nucleotides. The gaps that remain are sealed by DNA ligase .
Because eukaryotic chromosomes are linear, one might expect that their replication would be more straightforward. As in prokaryotes, the eukaryotic DNA polymerase can add nucleotides only in the 5’ to 3’ direction. In the leading strand, synthesis continues until it reaches either the end of the chromosome or another replication fork progressing in the opposite direction. On the lagging strand, DNA is synthesized in short stretches, each of which is initiated by a separate primer. When the replication fork reaches the end of the linear chromosome, there is no place to make a primer for the DNA fragment to be copied at the end of the chromosome. These ends thus remain unpaired and, over time, they may get progressively shorter as cells continue to divide.
The ends of the linear chromosomes are known as telomere s and consist of noncoding repetitive sequences. The telomeres protect coding sequences from being lost as cells continue to divide. In humans, a six base-pair sequence, TTAGGG, is repeated 100 to 1000 times to form the telomere. The discovery of the enzyme telomerase ( Figure 11.9 ) clarified our understanding of how chromosome ends are maintained. Telomerase contains a catalytic part and a built-in RNA template. It attaches to the end of the chromosome, and complementary bases to the RNA template are added on the 3’ end of the DNA strand. Once the 3’ end of the lagging strand template is sufficiently elongated, DNA polymerase can add the nucleotides complementary to the ends of the chromosomes. In this way, the ends of the chromosomes are replicated. In humans, telomerase is typically active in germ cells and adult stem cells; it is not active in adult somatic cells and may be associated with the aging of these cells. Eukaryotic microbes including fungi and protozoans also produce telomerase to maintain chromosomal integrity. For her discovery of telomerase and its action, Elizabeth Blackburn (1948–) received the Nobel Prize for Medicine or Physiology in 2009.
Link to Learning
This animation compares the process of prokaryotic and eukaryotic DNA replication.
- How does the origin of replication differ between eukaryotes and prokaryotes?
- What polymerase enzymes are responsible for DNA synthesis during eukaryotic replication?
- What is found at the ends of the chromosomes in eukaryotes and why?
DNA Replication of Extrachromosomal Elements: Plasmids and Viruses
To copy their nucleic acids, plasmids and viruses frequently use variations on the pattern of DNA replication described for prokaryote genomes. For more information on the wide range of viral replication strategies, see The Viral Life Cycle .
Rolling Circle Replication
Whereas many bacterial plasmids (see Unique Characteristics of Prokaryotic Cells ) replicate by a process similar to that used to copy the bacterial chromosome, other plasmids, several bacteriophages , and some viruses of eukaryotes use rolling circle replication ( Figure 11.10 ). The circular nature of plasmids and the circularization of some viral genomes on infection make this possible. Rolling circle replication begins with the enzymatic nicking of one strand of the double-stranded circular molecule at the double-stranded origin (dso) site . In bacteria, DNA polymerase III binds to the 3’-OH group of the nicked strand and begins to unidirectionally replicate the DNA using the un-nicked strand as a template, displacing the nicked strand as it does so. Completion of DNA replication at the site of the original nick results in full displacement of the nicked strand, which may then recircularize into a single-stranded DNA molecule. RNA primase then synthesizes a primer to initiate DNA replication at the single-stranded origin (sso) site of the single-stranded DNA (ssDNA) molecule, resulting in a double-stranded DNA (dsDNA) molecule identical to the other circular DNA molecule.
- Is there a lagging strand in rolling circle replication? Why or why not?
As an Amazon Associate we earn from qualifying purchases.
This book may not be used in the training of large language models or otherwise be ingested into large language models or generative AI offerings without OpenStax's permission.
Want to cite, share, or modify this book? This book uses the Creative Commons Attribution License and you must attribute OpenStax.
Access for free at https://openstax.org/books/microbiology/pages/1-introduction
- Authors: Nina Parker, Mark Schneegurt, Anh-Hue Thi Tu, Philip Lister, Brian M. Forster
- Publisher/website: OpenStax
- Book title: Microbiology
- Publication date: Nov 1, 2016
- Location: Houston, Texas
- Book URL: https://openstax.org/books/microbiology/pages/1-introduction
- Section URL: https://openstax.org/books/microbiology/pages/11-2-dna-replication
© Jan 10, 2024 OpenStax. Textbook content produced by OpenStax is licensed under a Creative Commons Attribution License . The OpenStax name, OpenStax logo, OpenStax book covers, OpenStax CNX name, and OpenStax CNX logo are not subject to the Creative Commons license and may not be reproduced without the prior and express written consent of Rice University.
An official website of the United States government
The .gov means it's official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you're on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
- Browse Titles
NCBI Bookshelf. A service of the National Library of Medicine, National Institutes of Health.
StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2024 Jan-.

StatPearls [Internet].
Biochemistry, dna replication.
Raheel Chaudhry ; Karam Khaddour .
Affiliations
Last Update: May 1, 2023 .
- Introduction
The existence of cell division implies that there is a mechanism that replicates DNA and supplies identical copies for the daughter cells while still maintaining an accurate representation of the genome. This mechanism, known as DNA replication, occurs in all organisms and allows for genetic inheritance. It can occur in a short period, copying up to approximately ten to the 11th power (10^11) units of information in some cases. The process of replication is semi-conservative, meaning that each of the two DNA molecules formed from the process is made up of one, old, template strand and one newly formed strand. It also forms the basis of the expression of genetic information through protein synthesis. Considering DNA replication occurs rapidly, there are also various mechanisms to ensure correct replication and minimize errors. Cancers can arise from errors or mutations in DNA replication. [1] [2] [3]
- Molecular Level
The blueprints of life are coded within the nucleic acid DNA (and for some organisms RNA). The structure of DNA is complex is made up of nucleotides, consisting of the sugar deoxyribose, a nitrogenous base (either adenine, guanine, cytosine, or thymine), and a phosphate group. Nucleotides made up of purine nitrogenous bases (adenine and guanine) can only pair with nucleotides made up of pyrimidine bases (thymine and cytosine, respectively). This isomorphism allows for base pairs to be replaced by one another and not ruin the backbone of the DNA. The monomers of the opposing strands stay together due to hydrogen bonds, and the two strands together form a double helix. Each strand runs antiparallel, meaning in opposite directions, one from the 5’ => 3’, the other 3’ => 5’ (This numbering comes from the carbon atoms in the sugar, which are labeled 1’ => 5’; the phosphate and hydroxyl group are attached to the 5’ and 3’ carbons respectively, creating the directionality of the nucleotide and, therefore, the DNA strand). The molecule is considered to be flexible, and that can be partially explained by the poor directionality of the hydrogen bonds which allow for bending or stretching. This flexibility enables many different DNA conformations, with high and low degrees of bending.
DNA is held in the nucleus of eukaryotes and within the cell membrane in the nucleoid for prokaryotes. It wraps around histone proteins to create nucleosomes, which group together to form chromatin and condense to form chromosomes. Chromatin can be negatively supercoiled (underwinded, more straightened) or positively supercoiled (overwinded, less straightened); this has implications in the use of DNA, either facilitating the binding of proteins to the nucleic acid or hindering it. The looser the DNA, the easier it is for proteins to access it, and the easier it is for replication to occur. Prokaryotic chromosomes are circular and are usually smaller in number. Eukaryotic chromosomes are linear and usually larger in number. Enzymes that are critical in DNA replication include DNA polymerase, helicase, topoisomerase, nuclease, ligase, and telomerase. These will be further elaborated on in the following sections.
The function of DNA replication is multifold and essential to life as we know it. This biological process allows for the genetic blueprints of a cell to be passed on to daughter cells in cell division without loss of genetic information. Without replication, when the cell divides, the information would be split and only partially passed on. [4] [5] [6]
DNA replication also allows for protein synthesis, which is how genes are expressed. Protein synthesis begins with transcribing the specific gene, or section of DNA, which codes for the desired protein. Without replication, a gene could have a limited number of outputs and could transcribe a limited number of proteins.
The actual mechanism is brought by the interaction of a multitude of proteins and enzymes and occurs during the S phase. The single DNA strand is separated into two complementary strands. DNA replication is a semiconservative process, meaning that for every new pair there is one original strand and one new strand.
The origin of replication is a sequence of base pairs in the genome where DNA replication begins; these sequences tend to be high in AT content making for easier separation. AT bonds have the fewest hydrogen bonds, making them weaker. Eukaryotes have multiple origins of replication whereas prokaryotes have one. Replication begins when origin-binding proteins bind to the origin of replication on the DNA. Then the unwinding of the double helix proceeds by way of the enzyme, helicase, creating a replication fork. The replication fork is Y-shaped and is where the leading and lagging strands are formed. Helicase begins unwinding by breaking hydrogen bonds. Single-stranded binding proteins (SSBs) stabilize the unwound DNA, preventing it from forming into secondary structures; the secondary structures can prevent the continuation of the DNA polymerase.
Meanwhile, topoisomerases reduce the pressure on the winded portions and continue to open the DNA downstream to allow for elongation. They work by changing the amount of DNA coiling by adding negative supercoils or by unlinking DNA circles for prokaryotic DNA. Topoisomerase I relaxes the supercoil, and topoisomerase II adds negative supercoils. Topoisomerase is known as DNA gyrase in prokaryotes. Once the DNA strand is open, there needs to be a base for the DNA polymerase to bind and begin replication; this is provided by the primase. Primase adds short strands of RNA primers (9 to 12 pairs) onto the template to allow for DNA polymerase III or DNA polymerase alpha to bind and add nucleotides. DNA polymerase III (in prokaryotes) has 5’ => 3’ synthesis and 3’ => 5’ proofreading exonuclease. DNA polymerase alpha (in eukaryotes) is a complex that has the DNA primase which creates the RNA primer, and then the polymerase alpha itself elongates around 20 nucleotides and passes off to DNA polymerase epsilon or delta . DNA polymerase epsilon elongates and proofreads on the leading strand. DNA polymerase delta elongates and proofreads on the lagging strand. Because the strands run antiparallel, the polymerization mechanism is slightly different for both strands. The DNA polymerase runs in the 3’ => 5’ direction (therefore creating DNA in the 5’ => 3’ orientation), but only one DNA template strand, known as the leading strand, is in the proper orientation. Replication of the leading strand is simple because it is already 3’ => 5’ direction, the polymerase continuously adds complementary nucleotides to the primer towards the replication fork. For the lagging strand, which is 5’ => 3’, the new strand is synthesized in segments and discontinuously because the DNA polymerase can only read in the 3’ => 5’ direction. A new primer is added as the replication fork is further opened and the DNA polymerase delta builds the new DNA strand away from the replication fork, creating Okazaki fragments. Afterwards, another DNA polymerase replaces the RNA primers with DNA; this is done by DNA polymerase I in prokaryotes and DNA polymerase delta in eukaryotes. DNA ligase connects the fragments.
Termination
In prokaryotes, replication ends when the forks meet. In eukaryotes replication ends at telomere regions. Telomeres are regions at the end of chromosomes with repetitive nucleotides such as TTAGGG sequences. Shortening telomeres have been associated with cell aging and death.
DNA Proofreading
The DNA replication is not perfect and has mechanisms to ensure there are corrections. DNA polymerases make mistakes in 1 in 10^5 base pairs. Considering the length of the human genome is around 3 x 10^9 that is about 30,000 to 50,000 mistakes. However, with the exonuclease activity of the polymerases, the error rates are reduced to a few hundred. These errors are further corrected in the G2 phase of the cell cycle, reducing the total number of errors to less than 10.
Polymerase Chain Reaction
Polymerase Chain Reaction (PCR) is a method of rapidly amplifying DNA. It is used clinically to test for the presence of specific bacteria and viruses in patients with certain diseases. Nucleic acid amplification testing uses PCR and is used to diagnose chlamydia. Also, reverse transcriptase PCR is used to detect HIV. [7]
Southern Blot [8]
Southern blotting is done by separating DNA fragments through electrophoresis, and then transferred the DNA from the electrophoresis gel to a filter. The filter is then washed with radioactively labeled hybridization probes that target for the DNA sequence of interest. Once the probes bind, the filter is exposed to x-ray, and the probes marking the DNA of interest are visible. Southern blot can be useful clinically for detecting mutations in certain DNA sequences.
- Clinical Significance
Antibiotics
Many antibiotics antivirals interfere with DNA replication in prokaryotes to prevent the replication of bacterias and viruses and the progression of disease in patients. One example includes fluoroquinolones, which bind to DNA topoisomerases and inhibit their function and preventing replication. Trimethoprim and sulfonamides prevent the formation of DNA precursors such as purines and pyrimidine, preventing DNA formation. Acyclovir acts as a guanosine analog which is monophosphorylated by HSV thymidine kinase; this phosphorylation creates a triphosphate which inhibits viral DNA polymerase by chain termination.
Antitumor Medications
Antitumor medications also may interfere with DNA replication. Cytarabine used to treat leukemia is a pyrimidine analog that inhibits DNA polymerase. 5-Fluorouracil, used in treating colon cancer, decreases the production of thymidine which is a nucleotide needed for DNA production. Etoposide is an antitumor medication that inhibits topoisomerase II; it is used to treat solid tumors, leukemias, and lymphomas. Irinotecan inhibits topoisomerase I.
- Review Questions
- Access free multiple choice questions on this topic.
- Comment on this article.
Disclosure: Raheel Chaudhry declares no relevant financial relationships with ineligible companies.
Disclosure: Karam Khaddour declares no relevant financial relationships with ineligible companies.
This book is distributed under the terms of the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International (CC BY-NC-ND 4.0) ( http://creativecommons.org/licenses/by-nc-nd/4.0/ ), which permits others to distribute the work, provided that the article is not altered or used commercially. You are not required to obtain permission to distribute this article, provided that you credit the author and journal.
- Cite this Page Chaudhry R, Khaddour K. Biochemistry, DNA Replication. [Updated 2023 May 1]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2024 Jan-.
In this Page
Bulk download.
- Bulk download StatPearls data from FTP
Related information
- PMC PubMed Central citations
- PubMed Links to PubMed
Similar articles in PubMed
- Review Organization of DNA replication. [Cold Spring Harb Perspect Biol...] Review Organization of DNA replication. Chagin VO, Stear JH, Cardoso MC. Cold Spring Harb Perspect Biol. 2010 Apr; 2(4):a000737.
- Enzyme-Free Replication with Two or Four Bases. [Angew Chem Int Ed Engl. 2018] Enzyme-Free Replication with Two or Four Bases. Hänle E, Richert C. Angew Chem Int Ed Engl. 2018 Jul 16; 57(29):8911-8915. Epub 2018 Jun 19.
- Review Principles and concepts of DNA replication in bacteria, archaea, and eukarya. [Cold Spring Harb Perspect Biol...] Review Principles and concepts of DNA replication in bacteria, archaea, and eukarya. O'Donnell M, Langston L, Stillman B. Cold Spring Harb Perspect Biol. 2013 Jul 1; 5(7). Epub 2013 Jul 1.
- Review Replication-Coupled DNA Repair. [Mol Cell. 2019] Review Replication-Coupled DNA Repair. Cortez D. Mol Cell. 2019 Jun 6; 74(5):866-876.
- Rolling circle RNA synthesis catalyzed by RNA. [Elife. 2022] Rolling circle RNA synthesis catalyzed by RNA. Kristoffersen EL, Burman M, Noy A, Holliger P. Elife. 2022 Feb 2; 11. Epub 2022 Feb 2.
Recent Activity
- Biochemistry, DNA Replication - StatPearls Biochemistry, DNA Replication - StatPearls
Your browsing activity is empty.
Activity recording is turned off.
Turn recording back on
Connect with NLM
National Library of Medicine 8600 Rockville Pike Bethesda, MD 20894
Web Policies FOIA HHS Vulnerability Disclosure
Help Accessibility Careers
DNA Replication Steps and Process
UIG / Getty Images
- Cell Biology
- Weather & Climate
- B.A., Biology, Emory University
- A.S., Nursing, Chattahoochee Technical College
DNA replication is the process in which a cell makes an identical copy of its DNA. It is vital for cell growth, repair, and reproduction in organisms as it helps with the transmission of genetic information. The replication process follows several steps involving multiple proteins called replication enzymes and RNA, or ribonucleic acid.
In eukaryotic cells, such as animal cells and plant cells , DNA replication occurs in the S phase of the cell cycle . Before this phase, also known as the synthesis stage, the cell passes through a preparation phase to minimize the chances of errors or mutations being introduced into the new DNA strands.
Key Takeaways
- Deoxyribonucleic acid, commonly known as DNA, is a nucleic acid that has three main components: a deoxyribose sugar, a phosphate, and a nitrogenous base.
- DNA contains the genetic material for an organism, and it must be copied when a cell divides into daughter cells. The process that copies DNA is called replication.
- Replication involves the production of identical helices of DNA from one double-stranded molecule of DNA.
- Enzymes are vital to DNA replication since they catalyze very important steps in the process.
- The overall DNA replication process is extremely important for both cell growth and reproduction in organisms. It is also vital in the cell repair process.
What Is DNA and Why Does It Replicate?
DNA is the genetic material that defines every cell. Before a cell duplicates and is divided into new daughter cells through either mitosis or meiosis , biomolecules and organelles must be copied to be distributed among the cells. DNA, found within the nucleus , must be replicated to ensure each new cell receives the correct number of chromosomes .
DNA Structure
DNA or deoxyribonucleic acid is a type of molecule known as a nucleic acid . It consists of a five-carbon deoxyribose sugar, a phosphate, and a nitrogenous base. Double-stranded DNA consists of two spiral nucleic acid chains that are twisted into a double helix shape. This twisting allows DNA to be more compact. To fit within the nucleus, DNA is packed into tightly coiled structures called chromatin . Chromatin condenses to form chromosomes during cell division. Before DNA replication, the chromatin loosens, giving cell replication machinery access to the DNA strands.
Replication Preparation and Beginning
Science Photo Library / Getty Images
Step 1: Replication Fork Formation
Before DNA can be replicated, the double-stranded molecule must be “unzipped” into two single strands. DNA has four bases called adenine (A), thymine (T), cytosine (C), and guanine (G) that form pairs between the two strands. Adenine only pairs with thymine and cytosine only binds with guanine. To unwind DNA, these interactions between base pairs must be broken. This is performed by an enzyme known as DNA helicase. DNA helicase disrupts the hydrogen bonding between base pairs to separate the strands into a Y shape known as the replication fork. This area will be the template for replication to begin.
DNA is directional in both strands, signified by a 5' and 3' end. This notation signifies which side group is attached to the DNA backbone. The 5' end has a phosphate (P) group attached, while the 3' end has a hydroxyl (OH) group attached. This directionality is important for replication as it only progresses in the 5' to 3' direction. However, the replication fork is bi-directional; one strand is oriented in the 3' to 5' direction (leading strand) while the other is oriented 5' to 3' (lagging strand). The two sides are therefore replicated with two different processes to accommodate the directional difference.
Replication Begins
Step 2: primer binding.
The leading strand is the simplest to replicate. Once the DNA strands have been separated, a short piece of RNA called a primer binds to the 3' end of the strand. The primer always binds as the starting point for replication. Primers are generated by the enzyme DNA primase.
DNA Replication: Elongation
UIG / Getty Images
Step 3: Elongation
Enzymes known as DNA polymerases are responsible for creating the new strand by a process called elongation. There are five different known types of DNA polymerases in bacteria and human cells . In bacteria such as E. coli, polymerase III is the main replication enzyme, while polymerase I, II, IV, and V are responsible for error checking and repair. DNA polymerase III binds to the strand at the site of the primer and begins adding new base pairs complementary to the strand during replication. In eukaryotic cells, polymerases alpha, delta, and epsilon are the primary polymerases involved in DNA replication. Because replication proceeds in the 5' to 3' direction on the leading strand, the newly formed strand is continuous.
The lagging strand begins replication by binding with multiple primers. Each primer is only several bases apart. DNA polymerase then adds pieces of DNA, called Okazaki fragments, to the strand between primers. This process of replication is discontinuous as the newly created fragments are disjointed.
Step 4: Termination
Once both the continuous and discontinuous strands are formed, an enzyme called exonuclease removes all RNA primers from the original strands. These primers are then replaced with appropriate bases. Another exonuclease “proofreads” the newly formed DNA to check, remove, and replace any errors. Another enzyme called DNA ligase joins Okazaki fragments together forming a single unified strand. The ends of the linear DNA present a problem as DNA polymerase can only add nucleotides in the 5′ to 3′ direction. The ends of the parent strands consist of repeated DNA sequences called telomeres. Telomeres act as protective caps at the end of chromosomes to prevent nearby chromosomes from fusing. A special type of DNA polymerase enzyme called telomerase catalyzes the synthesis of telomere sequences at the ends of the DNA. Once completed, the parent strand and its complementary DNA strand coils into the familiar double helix shape. In the end, replication produces two DNA molecules, each with one strand from the parent molecule and one new strand.
Replication Enzymes
Cultura / Getty Images
DNA replication would not occur without enzymes that catalyze various steps in the process. Enzymes that participate in the eukaryotic DNA replication process include:
- DNA helicase: Unwinds and separates double-stranded DNA as it moves along the DNA. It forms the replication fork by breaking hydrogen bonds between nucleotide pairs in DNA.
- DNA primase: A type of RNA polymerase that generates RNA primers. Primers are short RNA molecules that act as templates for the starting point of DNA replication.
- DNA polymerases: Synthesize new DNA molecules by adding nucleotides to leading and lagging DNA strands.
- Topoisomerase or DNA Gyrase: Unwinds and rewinds DNA strands to prevent the DNA from becoming tangled or supercoiled.
- Exonucleases: Group of enzymes that remove nucleotide bases from the end of a DNA chain.
- DNA ligase: Joins DNA fragments together by forming phosphodiester bonds between nucleotides.
DNA Replication Summary
Francis Leroy / Getty Images
DNA replication is the production of identical DNA helices from a single double-stranded DNA molecule. Each molecule consists of a strand from the original molecule and a newly formed strand. Prior to replication, the DNA uncoils and strands separate. A replication fork is formed which serves as a template for replication. Primers bind to the DNA and DNA polymerases add new nucleotide sequences in the 5′ to 3′ direction.
This addition is continuous in the leading strand and fragmented in the lagging strand. Once elongation of the DNA strands is complete, the strands are checked for errors, repairs are made, and telomere sequences are added to the ends of the DNA.
- Understanding the Double-Helix Structure of DNA
- DNA Definition: Shape, Replication, and Mutation
- Steps of Transcription From DNA to RNA
- An Introduction to DNA Transcription
- How Polymerase Chain Reaction Works to Amplify Genes
- What Is Polymerase Chain Reaction (PCR)?
- How Do Restriction Enzymes Cut DNA Sequences?
- Learn About Nucleic Acids and Their Function
- Nucleic Acids - Structure and Function
- DNA Sequencing Methods
- Learn How Virus Replication Occurs
- What Is RNA?
- Biology Prefixes and Suffixes: -ase
- What Are Restriction Enzymes?
- siRNA and How It Is Used
- What is Chromatin's Structure and Function?
- Reference Manager
- Simple TEXT file
People also looked at
Editorial article, editorial: the dna replication machinery as therapeutic targets.

- 1 New England Biolabs, Inc., Ipswich, MA, United States
- 2 Biomolecular Labeling Laboratory, Institute for Bioscience and Biotechnology Research, Rockville, MD, United States
- 3 National Institute of Standards and Technology, Rockville, MD, United States
Editorial on the Research Topic The DNA Replication Machinery as Therapeutic Targets
Chromosomal DNA replication is a process conserved through all domains of life. For accurate and efficient duplication of the genetic information, DNA replication components must work together in a highly regulated and coordinated fashion. Due to a central role in cell proliferation, the DNA replication machinery is an attractive therapeutic target for treating bacterial and viral infections, autoimmune disorders, and cancer. This eBook, entitled “The DNA Replication Machinery as Therapeutic Targets” explores how DNA replication factors serve as targets for new generations of therapies.
In all organisms, the chromosomal replication process starts at a specific chromosomal region called an origin of replication, at which origin-binding proteins (OBP) bind and locally unwind the duplex DNA. Additional proteins interact with the OBP-DNA complex and are responsible for the assembly of the DNA helicase around the DNA. Once assembled, the helicase unwinds the duplex in an ATP-dependent manner and forms the initial replication bubble. The exposed short regions of single-stranded DNA (ssDNA) at the replication bubble are coated with ssDNA-binding protein (SSB). DNA primase, DNA polymerases, and the rest of the replication machinery are recruited to the SSB-ssDNA complex to initiate bidirectional DNA synthesis. Due to the antiparallel nature of duplex DNA, and the unidirectionality of DNA polymerases, one strand of the chromosome is synthesized continuously (leading strand) while the other is copied discontinuously (lagging strand) as a series of Okazaki fragments ( O'Donnell et al., 2013 ; Kelman and Kelman, 2014 ; Kunkel and Burgers, 2017 ). Although these processes are fundamentally conserved in the three domains: archaea, bacteria and eukarya; as well as viruses and bacteriophages, the proteins, and complexes involved differ ( Makarova and Koonin, 2013 ).
Because the replication machinery is composed of a variety of core proteins and regulatory factors, disruption of any of the proteins involved will inhibit the replication process and/or its efficiency, and lead to replication stress. Replication stress is induced by endogenous factors such as dNTP depletion, DNA secondary structures or crosslinks, or by exogenous chemotherapies that damage DNA, such as cisplatin ( Kitao et al., 2018 ). In human cells, replication stress occurs when DNA polymerases uncouples from the replisome and lags behind helicase unwinding ( Zeman and Cimprich, 2014 ). As a result, long stretches of ssDNA are exposed at the replication fork. Replication protein A (RPA, the eukaryotic SSB) binds to the extended stretches of ssDNA, depleting free RPA in the cell and causing replication fork collapse (which may lead to DNA breakage and cell death). Therefore, inducing replication stress by disrupting the replication machinery or its regulation are an attractive strategies for drug design ( O'Connor, 2015 ; Forment and O'Connor, 2018 ).
Each chapter in this eBook highlights a different therapeutic strategy to effectively target DNA replication. One approach to halt replication is to inhibit DNA polymerases via binding of small molecules to the active site. The three replicative DNA polymerases responsible for the duplication of chromosomal DNA in eukarya, DNA polymerases α, δ and ε (Pol α, Pol δ, and Pol ε), all belong to family B DNA polymerases. In order to develop nucleotide analogs as drugs, one needs to understand how DNA polymerases incorporate natural and modified nucleotides into DNA. The contribution by Daimon et al. adds to our understanding of the structure and function relationships of family B polymerases by solving the structure of two members of this family from Aeropyrum pernix ( Daimon et al. ). The incorporation of various classes of modified nucleotides and nucleotide terminators by another family B DNA polymerase from archaea is described by a review contributed by Gardner et al . These archaeal polymerases share sequence similarity with the eukaryotic enzymes and thus can serve as good model systems for the more complex eukaryotic replication ( Makarova et al., 2014 ).
In addition, the use of nucleotide analogs to treat human autoimmune disorders and cancer is summarized in a contribution by Berdis . Young explores mitochondrial replication and how incorporation of certain nucleoside chain terminator inhibitors by Pol γ (the mitochondrial-specific polymerase) can lead to unintended toxicity by shutting down mitochondrial genome replication ( Young ). In addition, certain classes of compounds target Pol γ in cancer cells to inhibit mitochondrial replication with the potential to induce tumor cell death ( Young ).
Despite a central role in copying the chromosome, the inherent processivity of DNA polymerases is low, and only a few nucleotides are incorporated at a time. However, in the replication complex, high processivity DNA synthesis is conferred by a ring-shaped protein, referred to as the DNA polymerase sliding clamp, that encircles DNA and tethers the polymerase catalytic unit to the DNA for processive DNA synthesis ( Indiani and O'Donnell, 2006 ). The sliding clamps cannot assemble themselves around the DNA and require an additional clamp loader complex that assembles the clamp around duplex DNA in an ATP-dependent manner. In addition to interacting with the polymerase, the sliding clamps of bacteria and eukarya also interact with dozens of other proteins involved in DNA replication, repair, and cell cycle progression ( Vivona and Kelman, 2003 ). Therefore, inhibitors of the bacterial and eukaryal sliding clamps are being developed as anti-cancer and anti-bacterial drugs ( Georgescu et al., 2008 ). The current knowledge on the development of sliding clamps inhibitors and their possible use as therapeutic agents is summarized in a review contribution by Altieri and Kelman .
Another key enzyme for cellular replication is the DNA helicase, the enzyme responsible for unwinding double-stranded DNA ahead of the replisome ( Sakakibara et al., 2009 ). The contribution by Datta and Brosh describes the current state of the art in designing helicase inhibitors as anti-cancer drugs, and the issues surrounding the use of helicase inhibitors ( Datta and Brosh ). In eukarya, the replicative helicase is a complex of three components, the heterohexameric minichromosome maintenance (MCM), the tetrameric GINS complex and the Cdc45 protein. These form the CMG (Cdc45, MCM, GINS) complex ( Onesti and MacNeill, 2013 ; O'Donnell and Li, 2018 ). Due to the essential role of CMG in chromosome replication, it is a prime target for anti-cancer drugs. The current efforts in the development of CMG inhibitors as anti-cancer drugs are summarized in a contribution by Seo and Kang . Instead of directly inhibiting an enzyme activity (such as DNA polymerase), another strategy is to deplete activity by downregulating gene expression or deregulating protein activity via the ubiquitination pathway ( Jang et al .).
In addition, while some drugs are effective on their own, in other cases multiple replication factors can be targeted simultaneously to disrupt multiple pathways and lead to more efficient and effective treatment strategies. Mycobacterium tuberculosis is a pathogenic bacterium that is the etiological agent of tuberculosis (TB), which kills more than a million people a year ( Bañuls et al., 2015 ). Reiche and coauthors summarize the current state of the development of drugs against the M. tuberculosis replication machinery, including drugs targeting the polymerase (Pol III), the sliding clamp, clamp loader, and other replication proteins ( Reiche et al. ). In addition, Reiche et al. demonstrate the increased effectiveness of a combination M. tuberculosis antibiotic strategy that depletes dNTP pools while inhibiting DNA polymerase activity ( Reiche et al. ). Another example of a combination strategy is the inhibition of DNA polymerase synthesis with nucleotide inhibitors in combination with DNA damaging agents to create DNA lesion that stall synthesis ( Berdis ).
Remaining Challenges and Future Opportunities
Despite effective inhibitors of DNA replication proteins, eventual resistance to these inhibitors leads to tumor recurrence and remains a challenge for long-term therapeutic efficacy. Therefore, it will be important to continue to study molecular mechanisms of tumor resistance to DNA replication inhibitors. For example, DNA polymerase mutants that effectively remove nucleotide chain terminators can lead to drug resistance, as can upregulation of lesion bypass DNA polymerases ( Berdis ).
We anticipate that knowledge of DNA replication protein expression, regulation, and biochemical properties will continue to address these challenges and accelerate the development of novel strategies for effective treatment. To reach potential as therapeutic targets, more high-resolution structural information is needed for all replisome proteins and complexes to understand important replisome active site architectures and protein interactions. High resolution replisome structures will enable models for docking small molecules to inhibit enzyme activities and disrupt essential replisome interactions.
Finally, new molecular tools will accelerate identification of new DNA replication drugs targets. CRISPR-Cas9 genome engineering tools have revolutionized many scientific disciplines and offer a powerful method to alter genes by either modifying gene sequence or introducing insertions or deletions to knock out gene function ( Doudna and Charpentier, 2014 ). Genome-wide CRISPR-Cas9 screens aim to disrupt all or a subset of genes in an organism to identify important genes in a pathway ( Sánchez-Rivera and Jacks, 2015 ; Peters et al., 2016 ). CRISPR-Cas9 genome-wide screens can be adapted to identify novel factors that confer either resistance or sensitivity to DNA replication inhibitors. Knowledge of these factors may inform future therapeutic strategies to design new drug classes or enhance the efficacy of current therapies.
Author Contributions
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Conflict of Interest Statement
AG is employed and funded by New England Biolabs, Inc., a manufacturer and vendor of molecular biology reagents, including DNA replication and repair enzymes. This affiliation does not affect the author's impartiality, objectivity of data generation or its interpretation, adherence to journal standards and policies or availability of data.
The remaining author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Bañuls, A. L., Sanou, A., Anh, N. T., and Godreuil, S. (2015). Mycobacterium tuberculosis: ecology and evolution of a human bacterium. J. Med. Microbiol. 64, 1261–1269. doi: 10.1099/jmm.0.000171
PubMed Abstract | CrossRef Full Text | Google Scholar
Doudna, J. A., and Charpentier, E. (2014). Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096. doi: 10.1126/science.1258096
Forment, J. V., and O'Connor, M. J. (2018). Targeting the replication stress response in cancer. Pharmacol. Ther. 188, 155–167. doi: 10.1016/j.pharmthera.2018.03.005
Georgescu, R. E., Yurieva, O., Kim, S. S., Kuriyan, J., Kong, X. P., and O'Donnell, M. (2008). Structure of a small-molecule inhibitor of a DNA polymerase sliding clamp. Proc. Natl. Acad. Sci. U.S.A. 105, 11116–11121. doi: 10.1073/pnas.0804754105
Indiani, C., and O'Donnell, M. (2006). The replication clamp-loading machine at work in the three domains of life. Nat. Rev. Mol. Cell Biol. 7, 751–761. doi: 10.1038/nrm2022
Kelman, L. M., and Kelman, Z. (2014). Archaeal DNA replication. Annu. Rev. Genet. 48, 71–97. doi: 10.1146/annurev-genet-120213-092148
Kitao, H., Iimori, M., Kataoka, Y., Wakasa, T., Tokunaga, E., Saeki, H., et al. (2018). DNA replication stress and cancer chemotherapy. Cancer Sci. 109, 264–271. doi: 10.1111/cas.13455
Kunkel, T. A., and Burgers, P. M. J. (2017). Arranging eukaryotic nuclear DNA polymerases for replication: specific interactions with accessory proteins arrange Pols alpha, delta, and in the replisome for leading-strand and lagging-strand DNA replication. Bioessays 39:1700070. doi: 10.1002/bies.201700070
Makarova, K. S., and Koonin, E. V. (2013). Archaeology of eukaryotic DNA replication. Cold Spring Harb. Perspect. Biol. 5:a012963. doi: 10.1101/cshperspect.a012963
Makarova, K. S., Krupovic, M., and Koonin, E. V. (2014). Evolution of replicative DNA polymerases in archaea and their contributions to the eukaryotic replication machinery. Front. Microbiol. 5:354. doi: 10.3389/fmicb.2014.00354
O'Connor, M. J. (2015). Targeting the DNA damage response in cancer. Mol. Cell 60, 547–560. doi: 10.1016/j.molcel.2015.10.040
O'Donnell, M., Langston, L., and Stillman, B. (2013). Principles and concepts of DNA replication in bacteria, archaea, and eukarya. Cold Spring Harb. Perspect. Biol. 5:a010108. doi: 10.1101/cshperspect.a010108
O'Donnell, M. E., and Li, H. (2018). The ring-shaped hexameric helicases that function at DNA replication forks. Nat. Struct. Mol. Biol. 25, 122–130. doi: 10.1038/s41594-018-0024-x
Onesti, S., and MacNeill, S. A. (2013). Structure and evolutionary origins of the CMG complex. Chromosoma 122, 47–53. doi: 10.1007/s00412-013-0397-x
Peters, J. M., Colavin, A., Shi, H., Czarny, T. L., Larson, M. H., Wong, S., et al. (2016). A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506. doi: 10.1016/j.cell.2016.05.003
Sakakibara, N., Kelman, L. M., and Kelman, Z. (2009). Unwinding the structure and function of the archaeal MCM helicase. Mol. Microbiol. 72, 286–296. doi: 10.1111/j.1365-2958.2009.06663.x
Sánchez-Rivera, F. J., and Jacks, T. (2015). Applications of the CRISPR-Cas9 system in cancer biology. Nat. Rev. Cancer 15, 387–395. doi: 10.1038/nrc3950
Vivona, J. B., and Kelman, Z. (2003). The diverse spectrum of sliding clamp interacting proteins. FEBS Lett. 546, 167–172. doi: 10.1016/S0014-5793(03)00622-7
Zeman, M. K., and Cimprich, K. A. (2014). Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9. doi: 10.1038/ncb2897
Keywords: DNA replication, replication stress, DNA polymerase, helicase, therapeutic targets
Citation: Gardner AF and Kelman Z (2019) Editorial: The DNA Replication Machinery as Therapeutic Targets. Front. Mol. Biosci. 6:35. doi: 10.3389/fmolb.2019.00035
Received: 27 March 2019; Accepted: 02 May 2019; Published: 21 May 2019.
Reviewed by:
Copyright © 2019 Gardner and Kelman. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY) . The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Andrew F. Gardner, gardner@neb.com
This article is part of the Research Topic
The DNA Replication Machinery as Therapeutic Targets

IMAGES
VIDEO
COMMENTS
DNA replication is semiconservative, meaning that each strand in the DNA double helix acts as a template for the synthesis of a new, complementary strand. This process takes us from one starting molecule to two "daughter" molecules, with each newly formed double helix containing one new and one old strand.
Introduction. DNA synthesis occurs during the S phase of the cell cycle and is ensured by the replisome, a molecular machine made of a large number of proteins acting in a coordinated manner to synthesize DNA at many genomic locations, the replication origins 1.Replication origin activation in space and time (or replication program) is set by a sequence of events, starting already at the end ...
Accurate DNA replication is modulated by multiple replication‐associated proteins, which is fundamental to preserve genome stability. Abundant replication proteins are involved in tumorigenesis and development, implying these proteins act as therapeutic targets in clinical. Replication‐target cancer therapy emerges as the times require.
As discussed in Chapter 3, DNA replication is a semiconservative process in which each parental strand serves as a template for the synthesis of a new complementary daughter strand. The central enzyme involved is DNA polymerase, which catalyzes the joining of deoxyribonucleoside 5′-triphosphates (dNTPs) to form the growing DNA chain. However, DNA replication is much more complex than a ...
9.2: DNA Replication. When a cell divides, it is important that each daughter cell receives an identical copy of the DNA. This is accomplished by the process of DNA replication. The replication of DNA occurs during the synthesis phase, or S phase, of the cell cycle, before the cell enters mitosis or meiosis. The elucidation of the structure of ...
DNA replication has been extremely well-studied in prokaryotes, primarily because of the small size of the genome and large number of variants available. Escherichia coli has 4.6 million base pairs in a single circular chromosome, and all of it gets replicated in approximately 42 minutes, starting from a single origin of replication and ...
As previously mentioned, the location at which a DNA strand begins to unwind into two separate single strands is known as the origin of replication.As shown in Figure 1, when the double helix ...
DNA replication is the biological process by which an exact copy of a deoxyribonucleic acid (DNA) molecule is created and it is the basis for biological inheritance. Each of the two strands of the ...
DNA replication is the process of making an identical copy of the genetic material within each cell (Alberts et al. 2007; DePamphilis 2006).In eukaryotes, DNA replication takes place during a defined period of the cell cycle, called S (for synthesis) phase.DNA replication must be carried out with great precision every time the cell divides, so that genetic information is preserved.
which replication generates a double helix in which both strands are newly synthesized. Ergo, DNA replication is semi-conservative. DNA is synthesized by the repetitive addition of nucleotides to the 3' end of the growing polynucleotide chain DNA is synthesized by an iterative process in which nucleotides are added
The replication fork is a structure that forms within the long helical DNA during DNA replication. It is produced by enzymes called helicases that break the hydrogen bonds that hold the DNA strands together in a helix. The resulting structure has two branching "prongs", each one made up of a single strand of DNA.
The impact of replication stress on replication dynamics and DNA damage in vertebrate cells. Recent studies have provided insights into the sources of endogenous replication stress, which can ...
Figure 1: DNA Replication Process. Unwinding of DNA at the origin and synthesis of new strands. results in replication forks growing bidir ectional from the origin. number of proteins are ...
The process is quite rapid and occurs with few errors. DNA replication uses a large number of proteins and enzymes (Table 11.2.1 11.2. 1 ). One of the key players is the enzyme DNA polymerase, also known as DNA pol. In bacteria, three main types of DNA polymerases are known: DNA pol I, DNA pol II, and DNA pol III.
Matthew Meselson (1930-) and Franklin Stahl (1929-) devised an experiment in 1958 to test which of these models correctly represents DNA replication (Figure 11.5).They grew E. coli for several generations in a medium containing a "heavy" isotope of nitrogen (15 N) that was incorporated into nitrogenous bases and, eventually, into the DNA. This labeled the parental DNA.
The existence of cell division implies that there is a mechanism that replicates DNA and supplies identical copies for the daughter cells while still maintaining an accurate representation of the genome. This mechanism, known as DNA replication, occurs in all organisms and allows for genetic inheritance. It can occur in a short period, copying up to approximately ten to the 11th power (10^11 ...
DNA has directionality that can run either 3′-5′ or 5′-3′ based off of the carbons in the sugar group. The two strands of DNA in the double helix must run opposite to each other in an anti-parallel fashion. Therefore, if the first strand starts at the 3′ end and finishes at the 5′ end, then the second strand must run opposite, starting at the 5′ end and finishing at the 3′ end.
Dispersive replication. In the dispersive model, DNA replication results in two DNA molecules that are mixtures, or "hybrids," of parental and daughter DNA. In this model, each individual strand is a patchwork of original and new DNA. Most biologists at the time would likely have put their money on the semi-conservative model.
The replication process follows several steps involving multiple proteins called replication enzymes and RNA, or ribonucleic acid. In eukaryotic cells, such as animal cells and plant cells, DNA replication occurs in the S phase of the cell cycle. Before this phase, also known as the synthesis stage, the cell passes through a preparation phase ...
Chromosomal DNA replication is a process conserved through all domains of life. For accurate and efficient duplication of the genetic information, DNA replication components must work together in a highly regulated and coordinated fashion. Due to a central role in cell proliferation, the DNA replication machinery is an attractive therapeutic target for treating bacterial and viral infections ...
Summary: DNA replication takes place in three major steps. Opening of the double-stranded helical structure of DNA and separation of the strands. Priming of the template strands. Assembly of the newly formed DNA segments. During the separation of DNA, the two strands uncoil at a specific site known as the origin.
1. Time of Replication: The process of DNA replication takes place during cell division. The DNA replication takes place during S sub stage of interphase. In prokaryotes, DNA replication is initiated before the end of the cell cycle. Eukaryotic cells can only initiate DNA replication at the beginning of S phase. 2.
what is the basic process of dna replication through pcr. Here's the best way to solve it. Powered by Chegg AI. The basic process of DNA replication through PCR i... View the full answer. Previous question Next question. Not the question you're looking for? Post any question and get expert help quickly.