Gibbs Sampling
- Reference work entry
- First Online: 01 January 2018
- Cite this reference work entry

- Bruce E. Trumbo 3 &
- Eric A. Suess 3
211 Accesses
One of a number of computational methods collectively known as Markov chain Monte Carlo (MCMC) methods. In simulating a Markov chain, Gibbs sampling can be viewed as a special case of the Metropolis-Hastings algorithm. In statistical practice, the terminology Gibbs sampling most often refers to MCMC computations based on conditional distributions for the purpose of drawing inferences in multiparameter Bayesian models
Consider the parameter θ of a probability model as a random variable with the prior density function p (θ). The choice of a prior distribution may be based on previous experience or personal opinion. Then Bayesian inference combines information in the observed data x with information provided by the prior distribution to obtain a posterior distribution p ( θ | x ). The parameter θ may be a vector.
The likelihood function p ( x | θ ) is defined (up to a constant multiple) as the joint density function of the data x , now viewed as a...
This is a preview of subscription content, log in via an institution to check access.

Access this chapter
- Available as PDF
- Read on any device
- Instant download
- Own it forever
- Available as EPUB and PDF
- Durable hardcover edition
- Dispatched in 3 to 5 business days
- Free shipping worldwide - see info
Tax calculation will be finalised at checkout
Purchases are for personal use only
Institutional subscriptions
Box GEP, Tiao G (1973) Bayesian inference and statistical analysis. Addison-Wesley, Reading
MATH Google Scholar
Chib S, Greenberg E (1994) Understanding the Metropolis-Hastings algorithm. Am Stat 49:327–335
Google Scholar
Cox DR, Miller HD (1965) The theory of stochastic processes. Wiley, New York, MATH
Davies OL (ed) (1957) Statistical methods in research and production, 3rd edn. Oliver and Boyd, Edinburgh, MATH
Diaconis P, Freedman D (1997) On Markov chains with continuous state spaces (Statistics Technical Report 501), University of California Berkeley Library. www.stat.berkeley.edu/tech-reports/501.pdf
Donders FC (1868) Die Schnelligkeit psychischer Prozesse, Archiv für Anatomie und Physiologie und wissenschaftliche Medizin, pp. 657–681. English edition: Donders FC (1969) On the speed of mental processes, Attention and performance II (trans: Koster WG). North Holland, Amsterdam
Gastwirth JL (1987) The statistical precision of medical screening procedures: applications to polygraph and AIDS antibody test data (including discussion). Stat Sci 2:213–238, MATHMathSciNet
Article MathSciNet MATH Google Scholar
Gelfand AE, Smith AFM (1990) Sampling-based approaches to calculating marginal densities. J Am Stat Assoc 85:398–409, MATHMathSciNet
Gelman A, Carlin JB, Stern HS, Rubin DB (2004) Bayesian data analysis, 2nd edn. Chapman & Hall/CRC, Boca Raton, MATH
Hastings C Jr (1955) Approximations for digital computers. Princeton University Press, Princeton, MATH
Book MATH Google Scholar
Lee PM (2004) Bayesian statistics: an introduction, 3rd edn. Hodder Arnold, London
Metropolis N, Rosenbluth AW, Rosenbluth MN, Teller AH, Teller E (1953) Equations of state calculations by fast computing machines. J Chem Phys 21(6):1087–1092
Article Google Scholar
Pagano M, Gauvreau K (2000) Principles of biostatistics, 2nd edn. Belmont, Duxbury
R Development Core Team (2012) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, www.R-project.org. ISBN 3-900051-07-0
Robert CP, Casella G (2004) Monte Carlo statistical methods, 2nd edn. Springer, New York, MATH
Snedecor GW, Cochran WG (1980) Statistical methods, 7th edn. Iowa State University Press, Ahmmmmes
Speigelhalter D, Thomas A, Best N, Lunn D (2011) BUGS: Bayesian inference using Gibbs sampling. MRC Biostatistics Unit, Cambridge
Suess EA, Trumbo BE (2010) Introduction to probability simulation and Gibbs sampling with R. Springer, New York
Book Google Scholar
Trumbo BE (2002) Learning statistics with real data. Duxbury Press, Belmont
Download references
Author information
Authors and affiliations.
Department of Statistics and Biostatistics, California State University, East Bay, Hayward, CA, USA
Bruce E. Trumbo & Eric A. Suess
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Bruce E. Trumbo .
Editor information
Editors and affiliations.
Department of Computer Science, University of Calgary, Calgary, AB, Canada
Reda Alhajj
Section Editor information
American University of Sharjah, Sharjah, United Arab Emirates
Suheil Khoury
Rights and permissions
Reprints and permissions
Copyright information
© 2018 Springer Science+Business Media LLC, part of Springer Nature
About this entry
Cite this entry.
Trumbo, B.E., Suess, E.A. (2018). Gibbs Sampling. In: Alhajj, R., Rokne, J. (eds) Encyclopedia of Social Network Analysis and Mining. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-7131-2_146
Download citation
DOI : https://doi.org/10.1007/978-1-4939-7131-2_146
Published : 12 June 2018
Publisher Name : Springer, New York, NY
Print ISBN : 978-1-4939-7130-5
Online ISBN : 978-1-4939-7131-2
eBook Packages : Computer Science Reference Module Computer Science and Engineering
Share this entry
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
Help | Advanced Search
Statistics > Computation
Title: on gibbs sampling for structured bayesian models discussion of paper by zanella and roberts.
Abstract: This article is a discussion of Zanella and Roberts' paper: Multilevel linear models, gibbs samplers and multigrid decompositions. We consider several extensions in which the multigrid decomposition would bring us interesting insights, including vector hierarchical models, linear mixed effects models and partial centering parametrizations.
Submission history
Access paper:.
- Other Formats
References & Citations
- Google Scholar
- Semantic Scholar
BibTeX formatted citation
Bibliographic and Citation Tools
Code, data and media associated with this article, recommenders and search tools.
- Institution
arXivLabs: experimental projects with community collaborators
arXivLabs is a framework that allows collaborators to develop and share new arXiv features directly on our website.
Both individuals and organizations that work with arXivLabs have embraced and accepted our values of openness, community, excellence, and user data privacy. arXiv is committed to these values and only works with partners that adhere to them.
Have an idea for a project that will add value for arXiv's community? Learn more about arXivLabs .
Academia.edu no longer supports Internet Explorer.
To browse Academia.edu and the wider internet faster and more securely, please take a few seconds to upgrade your browser .
- We're Hiring!
- Help Center
Gibbs sampling
- Most Cited Papers
- Most Downloaded Papers
- Newest Papers
- Save to Library
- Last »
- Markov Chain Monte Carlo Follow Following
- Music Segmentation Follow Following
- Computer Sc Follow Following
- Amostrador De Gibbs Follow Following
- Ontology of Music Follow Following
- Markov chains Follow Following
- Cadeias De Markov Monte Carlo Follow Following
- Engenharia Florestal Follow Following
- Bayesian statistics & modelling Follow Following
- Statistical Analysis Techniques for Geography and Environmental Science Follow Following
Enter the email address you signed up with and we'll email you a reset link.
- Academia.edu Publishing
- We're Hiring!
- Help Center
- Find new research papers in:
- Health Sciences
- Earth Sciences
- Cognitive Science
- Mathematics
- Computer Science
- Academia ©2024
- Python for Machine Learning
- Machine Learning with R
- Machine Learning Algorithms
- Math for Machine Learning
- Machine Learning Interview Questions
- ML Projects
- Deep Learning
- Computer vision
- Data Science
- Artificial Intelligence
What is Gibbs Sampling?
- What is Upsampling in MATLAB?
- Systematic Sampling in R
- Stratified Sampling in R
- What makes a Sampling Data Reliable?
- Systematic Sampling in Pandas
- Stratified Sampling in Pandas
- Probability sampling
- Nyquist Sampling Theorem
- Cluster Sampling in R
- What is a Memory Pool?
- SQL Random Sampling within Groups
- Cluster Sampling in Pandas
- Methods of Sampling
- Cluster Sampling
- What is n in Statistics?
- What exactly Spooling is all about?
- Up-sampling in MATLAB
- Reservoir Sampling
- Sampling Theory
In statistics and machine learning, Gibbs Sampling is a potent Markov Chain Monte Carlo (MCMC) technique that is frequently utilized for sampling from intricate, high-dimensional probability distributions. The foundational ideas, mathematical formulas, and algorithm of Gibbs Sampling are examined in this article.
Gibbs Sampling
Gibbs sampling is a type of Markov Chain Monte Carlo(MCMC) method in which we sample from a set of multivariate(having different variables) probability distributions, it is typically considered to be difficult to sample joint distribution from the multivariate distribution so we consider conditional probability. Instead of calculating joint probability p(x 1 , x 2 , x 3 …..x n ), we will be calculating conditionals p(x i |x 2 , x 3 ….x n ) for every variable. The concept behind Gibbs Sampling is that we can update the value of variable x i keeping other variables having the current value. The process of updating variables and keeping others fixed creates a Markov Chain, after several iterations the chain converges to the joint distribution. Rather than finding the sample from joint distribution, we will prefer to find samples from conditional distribution for multivariate probability distribution which then tends to give us the joint distribution sampling.
Markov Chain Monte Carlo Method
Let’s discuss about Markov Chain Monte Carlo (MCMC) method in a little detail so that we can get intuition about the Gibbs Sampling.
In a Markov Chain, the future state depends on the current state and is independent of the previous state. To get a sample from the joint distribution, a Markov chain is created such that its stationary state corresponds to the target distribution. Until the stationary state is reached the samples taken into account are considered to be burn in state.
Monte Carlo methods corresponds to the random sampling of distribution when direct sampling from the distribution is not feasible.
In MCMC a random sample chain is formed X i -> X i+1 with the help of transition probability (T(X i+1 |X i )) which is actually the probability that the sample state X i will switch to X i+1 which is an implication that X i+1 marks higher density than X i in the p(x). We need to iterate over the process of switching from one sample to another until a series of stationary samples are obtained which tells us about the final state of samples, after this state the samples are not going to change and we can even match the stationary state with the target sample.
Let’s say that we start the sampling from X 0 , therefore the sampling process goes on like a chain – X 0 -> X 1 -> X 2 -> ………. X m -> X m -> X m+2 -> …and so on.
Here lets say that the samples starts matching the target distribution from X m , so the steps taken to reach X m form X 1 to X m-1 is considered to be burn in phase. The burn in phase is required to reach the required target samples.
The stationary state is reached when the distribution satisfies the equation =
where T(y|x) is the transition probability from y -> x and T(x|y) is the transition probability from x -> y
For the sake of simplicity let us consider the distribution is among three variables X, Y, Z and we want to sample joint distribution p(X, Y, Z).
Gibbs Sampling Algorithm
- First initialize each variable present in the system by assigning values to them.
- Then select any variable from the system X i , the selection choice of variable is not important.
- Now sample a new value of X i from the conditional distribution given other variables are the same p(X i |X 1 , X 2 ,…,X i-1 ,X i+1 ,……X n ) and update the value of X i in the sample.
- Repeat the iteration for a number of iterations with one variable at a time and for all the variables present in the system.
- A Markov Chain is created in the process which converges to the target distribution, we have to discard the initial values which led us to the target sample i.e. we have to discard the burn in phase.
Psuedocode for Gibbs Sampling Algorithm
The variables are initialized either randomly or with predetermined beginning values at the start of the Gibbs Sampling process. After that, iteratively cycling through each variable, it updates its value according to the conditional distribution based on how the other variables are currently doing. Taking into account the values of the nearby variables, a sample is taken from the conditional distribution of each variable x i . The current state is then updated using this sampled value. After a significant number of repetitions, the Markov chain is able to explore the joint distribution and eventually converge to the target distribution. This procedure is repeated. High-dimensional probability spaces can be effectively navigated by Gibbs Sampling due to its sequential variable update and dependence on conditional distributions.
Gibbs Sampling Function
Gibbs sampling involves various mathematical functions and expressions. Here are some common ones:
- Conditional Probability : We can express the conditional probability as P(x | y) , we use the vertical bar (|) within a fraction: [ P(x | y) ]
- Random variables : We can represent random variables as [X, Y, Z].
Steps to implement Gibbs Sampling Algorithm
- First we need to understand the problem and know the conditional probability of the variables we need to sample. Let’s say we have three variables X, Y, Z and we want to find their joint probability.
- After that we need to consider initial values for X, Y and Z, let’s say they are X 0 , Y 0 , Z 0 .
- Now we find the value of X keeping the value of Y and Z constant as the current values and randomly decide if we should change the value of X to the new value or the initial value, if we decide to take the new value we must update X from X 0 to X 1 .
- In the same way we find the value of Y keeping the value of X and Z constant as the current values and randomly decide if we should change the value of Y to the new value or the initial value, if we decide to take the new value we must update Y from Y 0 to Y 1 .
- Again we find the value of Z keeping the value of X and Y constant as the current values and randomly decide if we should change the value of Z to the new value or the initial value, if we decide to take the new value we must update Z from Z 0 to Z 1 .
- We keep repeating the third, fourth and the fifth steps to get new values of X, Y, Z and store their set (X i , Y j , Z k ) which will correspond to new samples.
- After a period the samples stored will converge to the target joint probability, we must consider recording the start of the joint probability and ignore the steps which were required to reach that distribution as those steps are considered to be burn in phase.
We can repeat what we just did with 3 variable distribution with ‘n’ variable distribution keeping ‘n-1’ variables constant.
Example of Gibbs sampling
Suppose we have two variables(X, Y) which takes two values 0 and 1, we want to sample from their joint distribution p(X, Y) which is defined as follows:
p(X=0, Y=0) = 0.2
p(X=1, Y=0) = 0.3
p(X=0, Y=1) = 0.1
p(X=1, Y=1) = 0.4
Let’s follow the above instructions to find out the join probability:
Initialize X and Y, let’s say X=0 and Y=0
- p(X=0|Y=0) = p(X=0,Y=0)/p(Y=0) = 0.2 / (0.3 + 0.2) = 0.4
- p(X=1|Y=0) = 1 – p(X=0, Y=0) = 1 – 0.4 = 0.6
We can see that the p(X=1|Y=0) > p(X=0|Y=0), therefore let’s consider X = 1
- p(Y=0|X=1) = 0.3 / (0.3 + 0.4) = 0.429
- p(Y=1|X=1) = 0.4 / (0.3 + 0.4) = 0.571
We can see that the p(Y=1|X=1) > p(Y=0|X=1), therefore let’s consider Y = 1
Now we get new sample (X = 1, Y = 1) from initial (X = 0, Y = 0) sample.
Implementation of Gibbs Sampling
Lets use real world dataset to perform Gibbs sampling on it, for this example we will be using iris flower dataset which has 4 features namely Sepal Length in cm, Sepal Width in cm, Petal Length in cm, Petal Width in cm and a target column named “Species” which specifies which specie of flower the given flower information belongs to. We will be using three features out of the features column and sampling from them, the three features that we are going to consider are Sepal Length in cm, Sepal Width in cm and Petal Length in cm and we will be sampling using Gibbs sampling. Now that we have the knowledge about the dataset lets sample from it to get more insightful and concrete knowledge about Gibbs Sampling. The link to get the iris dataset is present here – link .
Here in this code we imported three libraries named random to generate random numbers, pandas to manipulate data and we used and alias as ‘pd’ for it and numpy to compute numerical operations and again we used an alias as ‘np’. After that we read the csv file named ‘Iris.csv’ which is provided above and stored it into a variable named df. Then we prepared our data for gibbs sampling by dropping the columns ‘Species’ and ‘PetalWidthCm’ from the dataframe by using the method Dataframe.drop() and stored the remaining dataframe into a variable named data_pd, we did this since we will be sampling from three features named ‘SepalLengthCm’, ‘SepalWidthCm’ and ‘PetalLengthCm’. After this step we converted the data_pd into numpy array by using .to_numpy() method, which converted data_pd into a numpy matrix.
After following the above steps we started to get to the Gibbs Sampling part of the code, first we initialized three variables initial_x, initial_y, initial_z randomly from the range of values of the three features respectively. Then we assigned value to num_iterations variable that is how many samples that are going to be generated and it is set to 20, the stage to apply Gibbs Sampling has come so we iterate num_iterations time taking conditional of the three variables one at a time and changing the mean to new mean with standard deviation equals one and assigning new values to variables from random gauss distribution to get the samples.
Finally we will be appending all the samples to an empty list named ‘samples’ and printing it.

Advantages and Disadvantages of Gibbs Sampling
- Gibbs Sampling is relatively easy to implement if we compare it with other MCMC methods like Metrapolis-Hastings since it requires straightforward conditional distribution.
- Proposals are always accepted in Gibbs Sampling unlike Metrapolis-Hastings where accept-reject proposals happen.
- With the knowledge of conditional distribution of variables we can easily get the joint distribution of all the variables, which might get complex if we try direct method.
Disadvantages
- Gibbs Sampling may not be useful in case of complex distribution with irregular shapes where conditional distributions are difficult to obtain.
- It can be very slow in convergence if the variables are highly correlated.
- Conditional dependencies becomes complex if there are higher dimensions resulting in inaccuracy.
Please Login to comment...
Similar reads.
- Geeks Premier League 2023
- Geeks Premier League
- Machine Learning
Improve your Coding Skills with Practice
What kind of Experience do you want to share?
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- 09 May 2024
Cubic millimetre of brain mapped in spectacular detail
- Carissa Wong
You can also search for this author in PubMed Google Scholar
Rendering based on electron-microscope data, showing the positions of neurons in a fragment of the brain cortex. Neurons are coloured according to size. Credit: Google Research & Lichtman Lab (Harvard University). Renderings by D. Berger (Harvard University)
Researchers have mapped a tiny piece of the human brain in astonishing detail. The resulting cell atlas, which was described today in Science 1 and is available online , reveals new patterns of connections between brain cells called neurons, as well as cells that wrap around themselves to form knots, and pairs of neurons that are almost mirror images of each other.
The 3D map covers a volume of about one cubic millimetre, one-millionth of a whole brain, and contains roughly 57,000 cells and 150 million synapses — the connections between neurons. It incorporates a colossal 1.4 petabytes of data. “It’s a little bit humbling,” says Viren Jain, a neuroscientist at Google in Mountain View, California, and a co-author of the paper. “How are we ever going to really come to terms with all this complexity?”
Slivers of brain
The brain fragment was taken from a 45-year-old woman when she underwent surgery to treat her epilepsy. It came from the cortex, a part of the brain involved in learning, problem-solving and processing sensory signals. The sample was immersed in preservatives and stained with heavy metals to make the cells easier to see. Neuroscientist Jeff Lichtman at Harvard University in Cambridge, Massachusetts, and his colleagues then cut the sample into around 5,000 slices — each just 34 nanometres thick — that could be imaged using electron microscopes.
Jain’s team then built artificial-intelligence models that were able to stitch the microscope images together to reconstruct the whole sample in 3D. “I remember this moment, going into the map and looking at one individual synapse from this woman’s brain, and then zooming out into these other millions of pixels,” says Jain. “It felt sort of spiritual.”
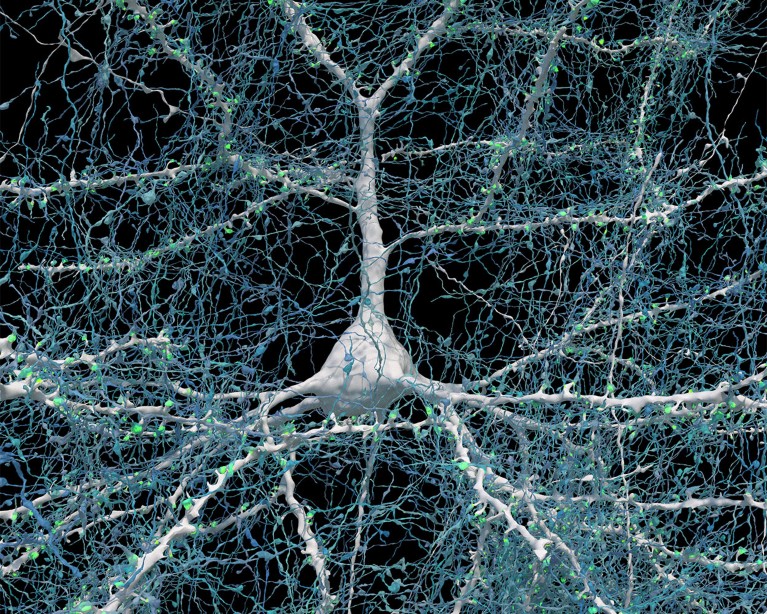
A single neuron (white) shown with 5,600 of the axons (blue) that connect to it. The synapses that make these connections are shown in green. Credit: Google Research & Lichtman Lab (Harvard University). Renderings by D. Berger (Harvard University)
When examining the model in detail, the researchers discovered unconventional neurons, including some that made up to 50 connections with each other. “In general, you would find a couple of connections at most between two neurons,” says Jain. Elsewhere, the model showed neurons with tendrils that formed knots around themselves. “Nobody had seen anything like this before,” Jain adds.
The team also found pairs of neurons that were near-perfect mirror images of each other. “We found two groups that would send their dendrites in two different directions, and sometimes there was a kind of mirror symmetry,” Jain says. It is unclear what role these features have in the brain.
Proofreaders needed
The map is so large that most of it has yet to be manually checked, and it could still contain errors created by the process of stitching so many images together. “Hundreds of cells have been ‘proofread’, but that’s obviously a few per cent of the 50,000 cells in there,” says Jain. He hopes that others will help to proofread parts of the map they are interested in. The team plans to produce similar maps of brain samples from other people — but a map of the entire brain is unlikely in the next few decades, he says.
“This paper is really the tour de force creation of a human cortex data set,” says Hongkui Zeng, director of the Allen Institute for Brain Science in Seattle. The vast amount of data that has been made freely accessible will “allow the community to look deeper into the micro-circuitry in the human cortex”, she adds.
Gaining a deeper understanding of how the cortex works could offer clues about how to treat some psychiatric and neurodegenerative diseases. “This map provides unprecedented details that can unveil new rules of neural connections and help to decipher the inner working of the human brain,” says Yongsoo Kim, a neuroscientist at Pennsylvania State University in Hershey.
doi: https://doi.org/10.1038/d41586-024-01387-9
Shapson-Coe, A. et al. Science 384 , eadk4858 (2024).
Article Google Scholar
Download references
Reprints and permissions
Related Articles
- Neuroscience

How does ChatGPT ‘think’? Psychology and neuroscience crack open AI large language models
News Feature 14 MAY 24
Brain-reading device is best yet at decoding ‘internal speech’
News 13 MAY 24
Retuning of hippocampal representations during sleep
Article 08 MAY 24
Found: the dial in the brain that controls the immune system
News 01 MAY 24
Assistant Scientist/Professor in Rare Disease Research, Sanford Research
Assistant Scientist/Professor in Rare Disease Research, Sanford Research Sanford Research invites applications for full-time faculty at the rank of...
Sioux Falls, South Dakota
Sanford Research
Postdoctoral Fellow - Boyi Gan lab
New postdoctoral positions are open in a cancer research laboratory located within The University of Texas MD Anderson Cancer Center. The lab curre...
Houston, Texas (US)
The University of Texas MD Anderson Cancer Center - Experimental Radiation Oncology
Assistant Professor
Tenure-track Assistant Professor position in the Cell and Molecular Physiology Department at Loyola University Chicago Stritch School of Medicine.
Maywood, Illinois
Loyola University of Chicago - Cell and Molecular Physiology Department
Chief Editor
We are looking for a Chief Editor to build and manage a team handling content at the interface of the physical and life sciences for the journal.
London or Berlin - hybrid working model.
Springer Nature Ltd
Junior and Senior Staff Scientists in microfluidics & optics
Seeking staff scientists with expertise in microfluidics or optics to support development of new technology to combat antimicrobial resistance
Boston, Massachusetts (US)
Harvard Medical School Systems Biology Department
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
MIT Technology Review
- Newsletters
Google helped make an exquisitely detailed map of a tiny piece of the human brain
A small brain sample was sliced into 5,000 pieces, and machine learning helped stitch it back together.
- Cassandra Willyard archive page
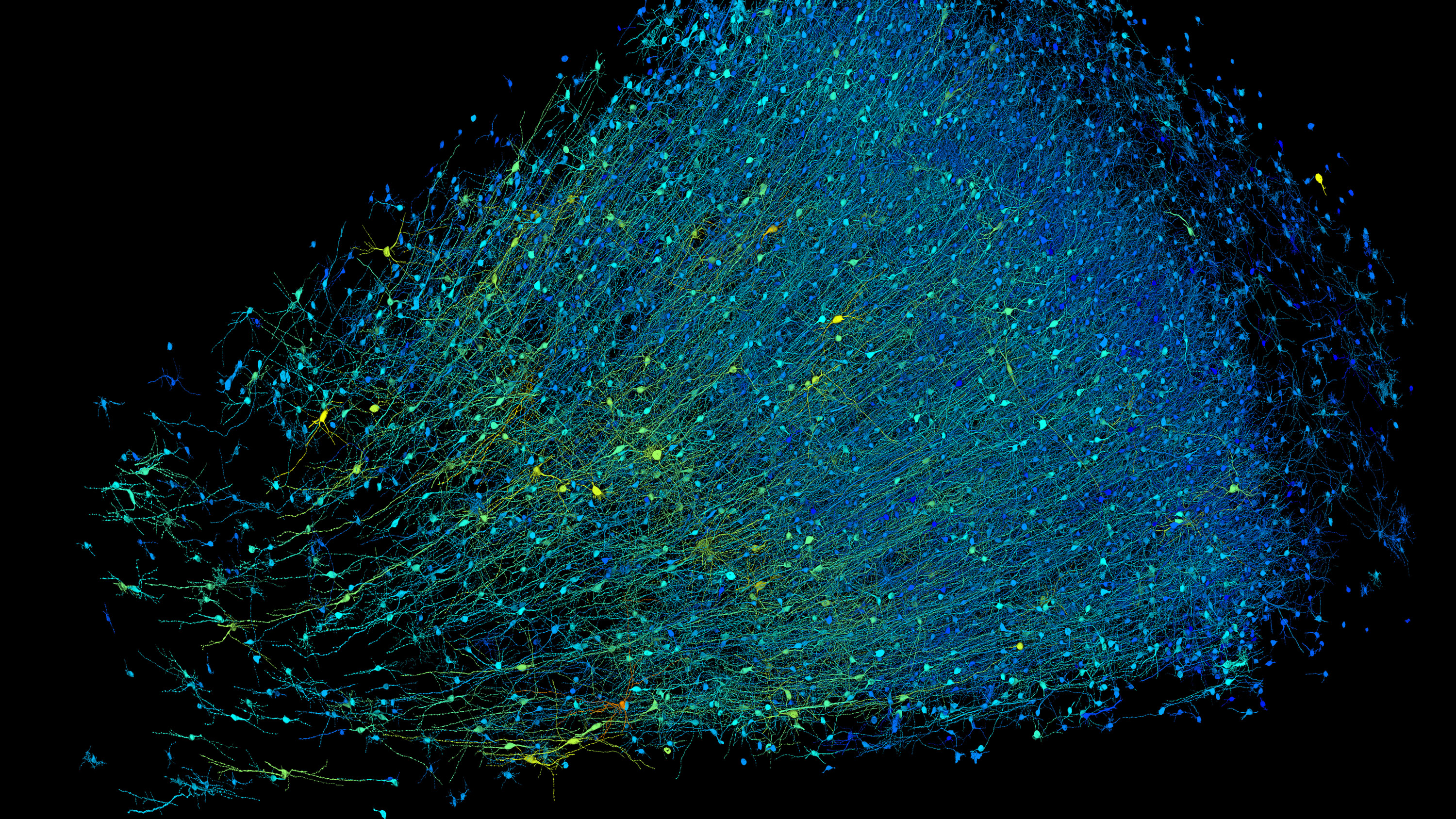
A team led by scientists from Harvard and Google has created a 3D, nanoscale-resolution map of a single cubic millimeter of the human brain. Although the map covers just a fraction of the organ—a whole brain is a million times larger—that piece contains roughly 57,000 cells, about 230 millimeters of blood vessels, and nearly 150 million synapses. It is currently the highest-resolution picture of the human brain ever created.
To make a map this finely detailed, the team had to cut the tissue sample into 5,000 slices and scan them with a high-speed electron microscope. Then they used a machine-learning model to help electronically stitch the slices back together and label the features. The raw data set alone took up 1.4 petabytes. “It’s probably the most computer-intensive work in all of neuroscience,” says Michael Hawrylycz, a computational neuroscientist at the Allen Institute for Brain Science, who was not involved in the research. “There is a Herculean amount of work involved.”
Many other brain atlases exist, but most provide much lower-resolution data. At the nanoscale, researchers can trace the brain’s wiring one neuron at a time to the synapses, the places where they connect. “To really understand how the human brain works, how it processes information, how it stores memories, we will ultimately need a map that’s at that resolution,” says Viren Jain, a senior research scientist at Google and coauthor on the paper, published in Science on May 9 . The data set itself and a preprint version of this paper were released in 2021 .
Brain atlases come in many forms. Some reveal how the cells are organized. Others cover gene expression. This one focuses on connections between cells, a field called “connectomics.” The outermost layer of the brain contains roughly 16 billion neurons that link up with each other to form trillions of connections. A single neuron might receive information from hundreds or even thousands of other neurons and send information to a similar number. That makes tracing these connections an exceedingly complex task, even in just a small piece of the brain..
To create this map, the team faced a number of hurdles. The first problem was finding a sample of brain tissue. The brain deteriorates quickly after death, so cadaver tissue doesn’t work. Instead, the team used a piece of tissue removed from a woman with epilepsy during brain surgery that was meant to help control her seizures.
Once the researchers had the sample, they had to carefully preserve it in resin so that it could be cut into slices, each about a thousandth the thickness of a human hair. Then they imaged the sections using a high-speed electron microscope designed specifically for this project.
Next came the computational challenge. “You have all of these wires traversing everywhere in three dimensions, making all kinds of different connections,” Jain says. The team at Google used a machine-learning model to stitch the slices back together, align each one with the next, color-code the wiring, and find the connections. This is harder than it might seem. “If you make a single mistake, then all of the connections attached to that wire are now incorrect,” Jain says.
“The ability to get this deep a reconstruction of any human brain sample is an important advance,” says Seth Ament, a neuroscientist at the University of Maryland. The map is “the closest to the ground truth that we can get right now.” But he also cautions that it’s a single brain specimen taken from a single individual.
The map, which is freely available at a web platform called Neuroglancer , is meant to be a resource other researchers can use to make their own discoveries. “Now anybody who’s interested in studying the human cortex in this level of detail can go into the data themselves. They can proofread certain structures to make sure everything is correct, and then publish their own findings,” Jain says. (The preprint has already been cited at least 136 times .)
The team has already identified some surprises. For example, some of the long tendrils that carry signals from one neuron to the next formed “whorls,” spots where they twirled around themselves. Axons typically form a single synapse to transmit information to the next cell. The team identified single axons that formed repeated connections—in some cases, 50 separate synapses. Why that might be isn’t yet clear, but the strong bonds could help facilitate very quick or strong reactions to certain stimuli, Jain says. “It’s a very simple finding about the organization of the human cortex,” he says. But “we didn’t know this before because we didn’t have maps at this resolution.”
The data set was full of surprises, says Jeff Lichtman, a neuroscientist at Harvard University who helped lead the research. “There were just so many things in it that were incompatible with what you would read in a textbook.” The researchers may not have explanations for what they’re seeing, but they have plenty of new questions: “That’s the way science moves forward.”
Biotechnology and health
How scientists traced a mysterious covid case back to six toilets.
When wastewater surveillance turns into a hunt for a single infected individual, the ethics get tricky.
An AI-driven “factory of drugs” claims to have hit a big milestone
Insilico is part of a wave of companies betting on AI as the "next amazing revolution" in biology
- Antonio Regalado archive page
The quest to legitimize longevity medicine
Longevity clinics offer a mix of services that largely cater to the wealthy. Now there’s a push to establish their work as a credible medical field.
- Jessica Hamzelou archive page
There is a new most expensive drug in the world. Price tag: $4.25 million
But will the latest gene therapy suffer the curse of the costliest drug?
Stay connected
Get the latest updates from mit technology review.
Discover special offers, top stories, upcoming events, and more.
Thank you for submitting your email!
It looks like something went wrong.
We’re having trouble saving your preferences. Try refreshing this page and updating them one more time. If you continue to get this message, reach out to us at [email protected] with a list of newsletters you’d like to receive.
Posted May 14, 2024
At 6:37 PM UTC
Wiley, a publishing company that’s more than 200 years old, is shuttering 19 journals today, the Wall Street Journal reports. Wiley has reportedly had to retract more than 11,300 papers recently “that appeared compromised” as generative AI makes it easier for paper mills to peddle fake research.
Lego Barad-dûr revealed: Sauron’s dark tower from The Lord of the Rings is $460
Openai releases gpt-4o, a faster model that’s free for all chatgpt users, apple ipad pro (2024) review: the best kind of overkill, the new apple ipad air is great — but it’s not the one to get, the washg1 is dyson’s first mop, more from science.
:format(webp)/cdn.vox-cdn.com/uploads/chorus_asset/file/25447321/2028780095.jpg)
US to raise tariffs on EVs, batteries, solar cells, and computer chips from China
:format(webp)/cdn.vox-cdn.com/uploads/chorus_asset/file/25447602/247109_Mars_Sample_Return_CVirginia_A.jpg)
The mission to retrieve a Mars sample is running into turbulence
:format(webp)/cdn.vox-cdn.com/uploads/chorus_asset/file/23144919/john_deere_4.jpg)
Solar storms made GPS tractors miss their mark at the worst time for farmers
:format(webp)/cdn.vox-cdn.com/uploads/chorus_asset/file/25442034/2118359708.jpg)
How to watch the possible aurora borealis this weekend

IMAGES
VIDEO
COMMENTS
The inadequate mixing of conventional Markov Chain Monte Carlo (MCMC) methods for multi-modal distributions presents a significant challenge in practical applications such as Bayesian inference and molecular dynamics. Addressing this, we propose Diffusive Gibbs Sampling (DiGS), an innovative family of sampling methods designed for effective sampling from distributions characterized by distant ...
Likelihood-free methods such as approximate Bayesian computation (ABC) have extended the reach of statistical inference to problems with computationally intractable likelihoods. Such approaches perform well for small-to-moderate dimensional problems, but suffer a curse of dimensionality in the number of model parameters. We introduce a likelihood-free approximate Gibbs sampler that naturally ...
Such dynamic sampling problems arise as a natural component of stochastic gradient methods for solving zero-sum games. We obtain our speedups by improving a Gibbs sampling subroutine developed in (van Apeldoorn & Gilyén,2019). We design a new dynamic quantum data structure perform- ing the necessary sampling in time Oe( 12), faster than the ...
This paper discusses how a variety of actuarial models can be implemented and ... We would like to thank the Greaves Undergraduate Research Program for ... Gibbs sampling in a similar area, however they had a focus on Whittaker-Henderson graduation. Additionally, Scollnik [10] performed a Bayesian analysis of a simultaneous equations model for ...
where the x i ∈ S and A ⊂ S is a possible set of states. Chains used in Gibbs sampling usually have continuous state spaces S. The index values i are often called steps or stages in time. Informally, the Markov Property is sometimes called "one-step dependence," but a sequence of independent random variables is trivially Markovian.A Markov chain can be fully specified by providing the ...
The Gibbs Sampler and Applications ... since the sample function will normalize theta anyway. Zhou, Qing/Monte Carlo Methods: Chapter 5 7 1.2. Stationary distribution and detail balance As a special case of the MH algorithm, the detail balance condition is satisfied
Under expanding domain asymptotics Davis and Borgman (1982) demonstrated that the distribution of each experimental variogram lag value converges to a Gaussian distribution. Fig. 3 (top row) shows the experimental variograms obtained for 50 realizations of GMRF-Gibbs sampler with practical range ρ = 50 and ν = 1 for fields of size 250 × 250 and 1 000 × 1 000.
In this paper, we will show that it is possible to use herding to generate samples from more complex unnormalized probability distributions. In particular, we introduce a de-terministic variant of the popular Gibbs sampling algorithm, which we refer to as herded Gibbs. While Gibbs relies on drawing samples from the full-conditionals at random ...
This article is a discussion of Zanella and Roberts' paper: Multilevel linear models, gibbs samplers and multigrid decompositions. We consider several extensions in which the multigrid decomposition would bring us interesting insights, including vector hierarchical models, linear mixed effects models and partial centering parametrizations.
Gibbs sampling is routinely used to sample truncated Gaussian distributions.These distributions naturally occur when associating latent Gaussian fields to category fields obtained by discrete simulation methods like multipoint, sequential indicator simulation and object-based simulation. The latent Gaussians are often used in data assimilation and history matching algorithms.
Search for more papers by this author. D. G. Clayton, D. G. Clayton. Medical Research Council Biostatistics Unit, Cambridge, UK ... Medical Research Council Biostatistics Unit, Institute of Public Health, University Forvie Site, ... We review applications of Gibbs sampling in medicine, involving longitudinal, spatial, covariate measurement and ...
2. Gibbs sampling 2.1. The Markov chain property 2.2. The Monte Carlo property 2.3. Checking the convergence 3. Some basic biology 4. Gibbs sampling for motif-finding 5. Gibbs sampling for biclustering gene expression data 6. Conclusion 1 Introduction Gibbs sampling is a technique to draw samples from a join distribution based
Chapter 5 - Gibbs Sampling. In this chapter, we will start describing Markov chain Monte Carlo methods. These methods are used to approximate high-dimensional expectations. (φ(X)) =. φ (x) (x) dx. and do not rely on independent samples from , or on the use of importance sampling. Instead, the samples are obtained by simulating a Markov chain ...
In statistics, Gibbs sampling or a Gibbs sampler is a Markov chain Monte Carlo (MCMC) algorithm for sampling from a specified multivariate probability distribution when direct sampling from the joint distribution is difficult, but sampling from the conditional distribution is more practical. This sequence can be used to approximate the joint distribution (e.g., to generate a histogram of the ...
An assessment of the impact of Gibbs sampling on the research community, on both statisticians and subject area scientists, and some thoughts on where the technology is headed and what needs to be done as the authors move into the next millennium are offered. During the course of the 1990s, the technology generally referred to as Markov chain Monte Carlo (MCMC) has revolutionized the way ...
Modelling Complexity: Applications of Gibbs Sampling in Medicine By W. R. GILKSt, D. G. CLAYTON, D. J. SPIEGELHALTER, N. G. BEST, A. J. McNEIL, L. D. SHARPLES and A. J. KIRBY Medical Research Council Biostatistics Unit, Cambridge, UK [Read before The Royal Statistical Society at a meeting on 'The Gibbs sampler and other Markov chain
For Gibbs sampling we have to be able to simulate from the following two conditional densities: 1) ̄ ; and 2) . We consider ̄ to be the ̄rst block x1, and Z ~ as the second block x2. Then by simulating ̄(t+1) and Z ~(t+1) j ̄(t+1); YT , we will get (at the limit) a draw from the joint distribution ̄; ZjYT ~ .
easy to sample from any of our f(x,θ) distributions directly (for example, sample θ from π(θ) and then sample x from f θ(x)). Further, we do not see how to extend present techniques to three or more component Gibbs samplers. 1.1 Notation and background Let (X,F) and (Θ,G) be measurable spaces equipped with σ-finite measures µ and ν ...
In this paper, based on the mixed Gibbs sampling algorithm, a Bayesian estimation procedure is proposed for a new Pareto-type distribution in the case of complete and type II censored samples. Simulation studies show that the proposed method is consistently superior to the maximize likelihood estimation in the context of small samples. Also, an analysis of some real data is provided to test ...
In this paper, motivated by a real dataset collected from a pharmaceutical company in China, we propose to use a novel predictive model that integrates a Bayesian framework with the Gibbs sampling ...
In this paper, a class of probability models for ranking data, the order-statistics models, is investigated. We extend the usual normal order-statistics model into one where the underlying random variables follow a multivariate normal distribution. Bayesian approach and the Gibbs sampling technique are used for parameter estimation.
The paper provides succinct guidelines for key sampling and data collection considerations in qualitative research involving interview studies. The importance of allowing time for immersion in a given community to become familiar with the context and population is discussed, as well as the practical constraints that sometimes operate against ...
In statistics and machine learning, Gibbs Sampling is a potent Markov Chain Monte Carlo (MCMC) technique that is frequently utilized for sampling from intricate, high-dimensional probability distributions. The foundational ideas, mathematical formulas, and algorithm of Gibbs Sampling are examined in this article.
The 3D map covers a volume of about one cubic millimetre, one-millionth of a whole brain, and contains roughly 57,000 cells and 150 million synapses — the connections between neurons. It ...
A team led by scientists from Harvard and Google has created a 3D, nanoscale-resolution map of a single cubic millimeter of the human brain. Although the map covers just a fraction of the organ ...
J. Justine Calma. A surge in fraudulent research papers is shutting down scientific journals. Wiley, a publishing company that's more than 200 years old, is shuttering 19 journals today, the ...