Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
- Dissertation
- Thesis & Dissertation Acknowledgements | Tips & Examples

Thesis & Dissertation Acknowledgements | Tips & Examples
Published on May 3, 2022 by Tegan George . Revised on July 18, 2023.
The acknowledgements section is your opportunity to thank those who have helped and supported you personally and professionally during your thesis or dissertation process.
Thesis or dissertation acknowledgements appear between your title page and abstract and should be no longer than one page.
In your acknowledgements, it’s okay to use a more informal style than is usually permitted in academic writing , as well as first-person pronouns . Acknowledgements are not considered part of the academic work itself, but rather your chance to write something more personal.
To get started, download our step-by-step template in the format of your choice below. We’ve also included sample sentence starters to help you construct your acknowledgments section from scratch.
Download Word doc Download Google doc
Instantly correct all language mistakes in your text
Upload your document to correct all your mistakes in minutes
Table of contents
Who to thank in your acknowledgements, how to write acknowledgements, acknowledgements section example, acknowledgements dos and don’ts, other interesting articles, frequently asked questions about the acknowledgements section.
Generally, there are two main categories of acknowledgements: professional and personal .
A good first step is to check your university’s guidelines, as they may have rules or preferences about the order, phrasing, or layout of acknowledgements. Some institutions prefer that you keep your acknowledgements strictly professional.
Regardless, it’s usually a good idea to place professional acknowledgements first, followed by any personal ones. You can then proceed by ranking who you’d like to thank from most formal to least.
- Chairs, supervisors, or defense committees
- Funding bodies
- Other academics (e.g., colleagues or cohort members)
- Editors or proofreaders
- Librarians, research/laboratory assistants, or study participants
- Family, friends, or pets
Typically, it’s only necessary to mention people who directly supported you during your thesis or dissertation. However, if you feel that someone like a high school physics teacher was a great inspiration on the path to your current research, feel free to include them as well.
Professional acknowledgements
It is crucial to avoid overlooking anyone who helped you professionally as you completed your thesis or dissertation. As a rule of thumb, anyone who directly contributed to your research process, from figuring out your dissertation topic to your final proofread, should be mentioned.
A few things to keep in mind include:
- Even if you feel your chair didn’t help you very much, you should still thank them first to avoid looking like you’re snubbing them.
- Be sure to follow academic conventions, using full names with titles where appropriate.
- If several members of a group or organization assisted you, mention the collective name only.
- Remember the ethical considerations around anonymized data. If you wish to protect someone’s privacy, use only their first name or a generic identifier (such as “the interviewees”)/
Personal acknowledgements
There is no need to mention every member of your family or friend group. However, if someone was particularly inspiring or supportive, you may wish to mention them specifically. Many people choose to thank parents, partners, children, friends, and even pets, but you can mention anyone who offered moral support or encouragement, or helped you in a tangible or intangible way.
Some students may wish to dedicate their dissertation to a deceased influential person in their personal life. In this case, it’s okay to mention them first, before any professional acknowledgements.
Don't submit your assignments before you do this
The academic proofreading tool has been trained on 1000s of academic texts. Making it the most accurate and reliable proofreading tool for students. Free citation check included.
Try for free
After you’ve compiled a list of who you’d like to thank, you can then sort your list into rank order. Separate everyone you listed into “major thanks,” “big thanks,” and “minor thanks” categories.
- “Major thanks” are given to people who your project would be impossible without. These are often predominantly professional acknowledgements, such as your advisor, chair, and committee, as well as any funders.
- “Big thanks” are an in-between, for those who helped you along the way or helped you grow intellectually, such as classmates, peers, or librarians.
- “Minor thanks” can be a catch-all for everyone else, especially those who offered moral support or encouragement. This can include personal acknowledgements, such as parents, partners, children, friends, or even pets.
How to phrase your acknowledgements
To avoid acknowledgements that sound repetitive or dull, consider changing up your phrasing. Here are some examples of common sentence starters you can use for each category.
| Major thanks | Big thanks | Minor thanks |
|---|---|---|
Note that you do not need to write any sort of conclusion or summary at the end. You can simply end the acknowledgements with your last thank you.
Here’s an example of how you can combine the different sentences to write your acknowledgements.
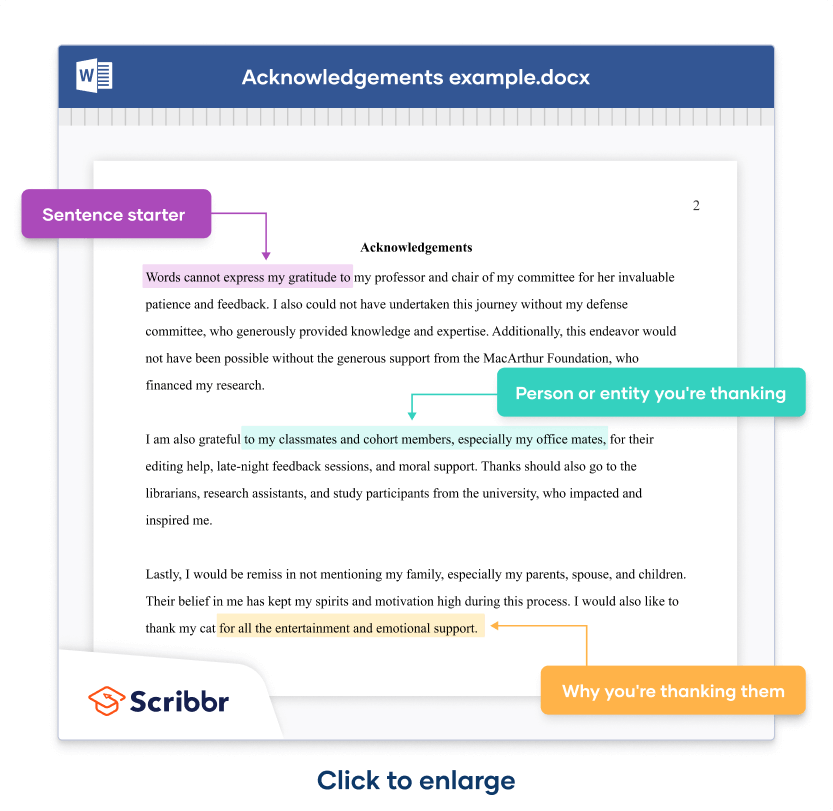
A simple construction consists of a sentence starter (in purple highlight ), followed by the person or entity mentioned (in green highlight ), followed by what you’re thanking them for (in yellow highlight .)
Acknowledgements
Words cannot express my gratitude to my professor and chair of my committee for her invaluable patience and feedback. I also could not have undertaken this journey without my defense committee, who generously provided knowledge and expertise. Additionally, this endeavor would not have been possible without the generous support from the MacArthur Foundation, who financed my research .
I am also grateful to my classmates and cohort members, especially my office mates, for their editing help, late-night feedback sessions, and moral support. Thanks should also go to the librarians, research assistants, and study participants from the university, who impacted and inspired me.
Lastly, I would be remiss in not mentioning my family, especially my parents, spouse, and children. Their belief in me has kept my spirits and motivation high during this process. I would also like to thank my cat for all the entertainment and emotional support.
- Write in first-person, professional language
- Thank your professional contacts first
- Include full names, titles, and roles of professional acknowledgements
- Include personal or intangible supporters, like friends, family, or even pets
- Mention funding bodies and what they funded
- Appropriately anonymize or group research participants or non-individual acknowledgments
Don’t:
- Use informal language or slang
- Go over one page in length
- Mention people who had only a peripheral or minor impact on your work
Here's why students love Scribbr's proofreading services
Discover proofreading & editing
If you want to know more about AI for academic writing, AI tools, or research bias, make sure to check out some of our other articles with explanations and examples or go directly to our tools!
Research bias
- Anchoring bias
- Halo effect
- The Baader–Meinhof phenomenon
- The placebo effect
- Nonresponse bias
- Deep learning
- Generative AI
- Machine learning
- Reinforcement learning
- Supervised vs. unsupervised learning
(AI) Tools
- Grammar Checker
- Paraphrasing Tool
- Text Summarizer
- AI Detector
- Plagiarism Checker
- Citation Generator
In the acknowledgements of your thesis or dissertation, you should first thank those who helped you academically or professionally, such as your supervisor, funders, and other academics.
Then you can include personal thanks to friends, family members, or anyone else who supported you during the process.
Yes, it’s important to thank your supervisor(s) in the acknowledgements section of your thesis or dissertation .
Even if you feel your supervisor did not contribute greatly to the final product, you must acknowledge them, if only for a very brief thank you. If you do not include your supervisor, it may be seen as a snub.
The acknowledgements are generally included at the very beginning of your thesis , directly after the title page and before the abstract .
In a thesis or dissertation, the acknowledgements should usually be no longer than one page. There is no minimum length.
You may acknowledge God in your dissertation acknowledgements , but be sure to follow academic convention by also thanking the members of academia, as well as family, colleagues, and friends who helped you.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the “Cite this Scribbr article” button to automatically add the citation to our free Citation Generator.
George, T. (2023, July 18). Thesis & Dissertation Acknowledgements | Tips & Examples. Scribbr. Retrieved June 24, 2024, from https://www.scribbr.com/dissertation/acknowledgements/
Is this article helpful?
Tegan George
Other students also liked, dissertation layout and formatting, thesis & dissertation title page | free templates & examples, how to write an abstract | steps & examples, get unlimited documents corrected.
✔ Free APA citation check included ✔ Unlimited document corrections ✔ Specialized in correcting academic texts
Acknowledgement in Research Paper – A Quick Guide [5 Examples]
The acknowledgement section in your research paper is where you thank those who have helped or supported you throughout your research and writing. It is a short section of 3-5 paragraphs or no more than 300 words you put on a page after the title page.
Saying thank you with style
How to write an acknowledgement: the complete guide for students, why should i include an acknowledgement in my research paper.
Acknowledging assistance and contributions from others can establish your integrity as a researcher. This will eventually make your work more credible.
What should be acknowledged about (aka thankful for)?
Who should be included in the acknowledgement of a research paper.
You can include everyone who helped you technically, intellectually, or financially (assistance with grants or monetary help) in the process of researching and writing your research paper. Except for your family and friends, you should always include the full names with the title of these individuals:
You can start with your professor or the individuals who supported you the most throughout the research. And then you can continue by thanking your institution and then the reviewer who reviewed your paper. Then you can thank your friends and families and any other individual who helped.
What is the tone of the acknowledgement in a research paper?
You should write your acknowledgement in formal language with complete sentences. It is appropriate to write in the first person (‘I’ for a single author or ‘we’ for two or more).
Writing an acknowledgement for research paper is one of the important parts of your project report. You need to thank everyone for helping you with your paper . Here are some examples of acknowledgement for your research paper.
Acknowledgement in Research Paper: Example 1
Acknowledgement in research paper: example 2, acknowledgement in research paper: example 3, acknowledgement in research paper: example 4, acknowledgement in research paper: example 5.
You can use these or try to create your own version for your project report. Also, you can use our auto acknowledgement generator tool to automatically generate acknowledgement for your project.
Where should I put the acknowledgement section?
How long is an acknowledgement in a research paper.
The acknowledgement section (usualy inserted as a page) of your research paper should consist of 3-5 paragraphs or no more than 300 words you put on a page after the title page.
Should I use the full names of family members in an acknowledgement?
You do not necessarily need to use the full name for your family and friends (it would sound pretty awkward to use the full name of your parent or spouse right?), you should always include the full names with the title for all other individuals in your acknowledgement.
Can I use “first person” in an acknowledgement?
What is an acknowledgement in academic writing?
More Definitions on Acknowledgement
“acknowledgement” vs “acknowledgment”… …what the hack.
Both “acknowledgement” and “acknowledgment” are used in the English-speaking world. However, acknowledgement with the “e” in the middle is more commonly used. It is up to 24.5 times more popular in the top 5 English-speaking countries in the world.
Other Popular Acknowledgement Examples
Others Acknowledgement to Funeral Attendees [5 Examples] Funeral Acknowledgement Templates (for Newspapers and Websites) Common Website Disclaimers to Protect Your Online Business Notary Acknowledgement [5 Examples]
Acknowledgement Examples for School/College Projects
How-to guides on academic writing and others, introduction to academic writing.
By O.P. Jindal Global University Duration: 16-hour Cost: FREE Gain an in-depth understanding of reading and writing as essential skills to conduct robust and critical research for your writing.
Writing in English at University
Academic english: writing specialization, 4 thoughts on “acknowledgement in research paper – a quick guide [5 examples]”.
This is great! Your samples really helped me in my research. Thank you and more power!
Leave a Comment Cancel Reply
Your email address will not be published. Required fields are marked *
Thesis Acknowledgements Examples
5 Examples For Your Inspiration
By: Derek Jansen (MBA) | Reviewers: Dr Eunice Rautenbach | May 2024
The acknowledgements section of your thesis or dissertation is an opportunity to say thanks to all the people who helped you along your research journey. In this post, we’ll share five thesis acknowledgement examples to provide you with some inspiration.
Overview: Acknowledgements Examples
- The acknowledgements 101
- Example: Formal and professional
- Example: Warm and personal
- Example: Challenge-centric
- Example: Institution-centric
- Example: Reflective and philosophical
- FREE Acknowledgements template
Acknowledgements 101: The Basics
The acknowledgements section in your thesis or dissertation is where you express gratitude to those who helped bring your project to fruition. This section is typically brief (a page or less) and less formal, but it’s crucial to thank the right individuals in the right order .
As a rule of thumb, you’ll usually begin with academic support : your supervisors, advisors, and faculty members. Next, you’ll acknowledge any funding bodies or sponsors that supported your research. You’ll then follow this with your intellectual contributors , such as colleagues and peers. Lastly, you’ll typically thank your personal support network , including family, friends, and even pets who offered emotional support during your studies.
As you can probably see, this order moves from the most formal acknowledgements to the least . Typically, your supervisor is mentioned first, due to their significant role in guiding and potentially evaluating your work. However, while this structure is recommended, it’s essential to adapt it based on any specific guidelines from your university. So, be sure to always check (and adhere to) any requirements or norms specific to your university.
With that groundwork laid, let’s look at a few dissertation and thesis acknowledgement examples . If you’d like more, check out our collection of dissertation examples here .
Need a helping hand?
Example #1: Formal and Professional
This acknowledgement formally expresses gratitude to academic mentors and peers, emphasising professional support and academic guidance.
I extend my deepest gratitude to my supervisor, Professor Jane Smith, for her unwavering support and insightful critiques throughout my research journey. Her deep commitment to academic excellence and meticulous attention to detail have significantly shaped this dissertation. I am equally thankful to the members of my thesis committee, Dr. John Doe and Dr. Emily White, for their constructive feedback and essential suggestions that enhanced the quality of my work.
My appreciation also goes to the faculty and staff in the Department of Biology at University College London, whose resources and assistance have been invaluable. I would also like to acknowledge my peers for their camaraderie and the stimulating discussions that inspired me throughout my academic journey. Their collective wisdom and encouragement have been a cornerstone of my research experience.
Finally, my sincere thanks to the technical staff whose expertise in managing laboratory equipment was crucial for my experiments. Their patience and readiness to assist at all times have left a profound impact on the completion of my project.
Example #2: Warm and Personal
This acknowledgement warmly credits the emotional and personal support received from family, friends and an approachable mentor during the research process.
This thesis is a reflection of the unwavering support and boundless love I received from my family and friends during this challenging academic pursuit. I owe an immense debt of gratitude to my parents, who nurtured my curiosity and supported my educational endeavours from the very beginning.
To my partner, Alex, your endless patience and understanding, especially during the most demanding phases of this research, have been my anchor. A heartfelt thank you to my supervisor, Dr. Mark Brown, whose mentorship extended beyond the academic realm into personal guidance, providing comfort and motivation during tough times.
I am also thankful for my friends, who provided both distractions when needed and encouragement when it seemed impossible to continue. The discussions and unwavering support from my peers at the university have enriched my research experience, making this journey not only possible but also enjoyable. Their presence and insights have been pivotal in navigating the complexities of my research topic.
Example #3: Inclusive of Challenges
This acknowledgement recognises the wide range of support received during unexpected personal and academic challenges, highlighting resilience and communal support.
The path to completing this dissertation has been fraught with both professional challenges and personal adversities. I am profoundly grateful for the enduring support of my supervisor, Dr. Lisa Green, whose steadfast belief in my capabilities and academic potential encouraged me to persevere even during my lowest moments. Her guidance was a beacon of light in times of doubt.
I must also express my deepest appreciation for my family, who stepped in not only with emotional reassurance, but also with critical financial support, when unexpected personal challenges arose. Their unconditional love and sacrifice have been the foundation of my resilience and success.
I am thankful, too, for the support services at the university, including the counselling centre, whose staff provided me with the tools to manage stress and maintain focus on my academic goals. Each of these individuals and institutions played a crucial role in my journey, reminding me that the pursuit of knowledge is not a solo expedition but a communal effort.
Example #4: Institution-Centric
This acknowledgement succinctly appreciates the financial, academic, and operational support provided by the university and its staff.
This dissertation would not have been possible without the generous financial support from the University of Edinburgh Scholarship Programme, which enabled me to focus fully on my research without financial burden. I am profoundly grateful to my advisor, Professor Richard Miles, for his sage advice, rigorous academic guidance, and the confidence he instilled in me. His expertise in the field of microbiology greatly enriched my work.
I would also like to thank the staff at the university library, whose assistance was indispensable. Their patience and willingness to help locate rare and essential resources facilitated a thorough and comprehensive literature review. Additionally, the administrative and technical staff, who often go unmentioned, provided necessary support that greatly enhanced my research experience. Their dedication and hard work create an environment conducive to academic success.
Lastly, my peers’ encouragement and the intellectual environment at the university have been vital in completing my research.
Example #5: Reflective and philosophical
This acknowledgement reflects on the philosophical growth and personal insights gained through the support of mentors, peers, and family, framing the dissertation as a journey of intellectual discovery.
Completing this thesis has been not only an academic challenge but also a profound journey of personal and philosophical growth.
I am immensely grateful to my mentor, Professor Sarah Johnson, for encouraging me to explore complex ideas and to challenge conventional wisdom. Her guidance helped me navigate the philosophical underpinnings of my research and deepened my analytical skills. My fellow students in the Philosophy Department provided a supportive and intellectually stimulating community. Our discussions extended beyond the classroom, offering new insights and perspectives that were crucial to my thesis.
I am also deeply thankful for my family, whose intellectual curiosity and spirited philosophical debates at home sparked my interest in philosophy from a young age. Their continuous support and belief in my academic pursuits have been instrumental in my success.
This dissertation reflects not only my work but also the collective support of everyone who has touched my life academically and personally. The journey has taught me the value of questioning and the importance of diverse perspectives in enriching our understanding of complex philosophical issues.

FREE Acknowledgements Template
To help fast-track your writing process, we’ve created a free, fully editable template . This template covers all the necessary content for a generic thesis or dissertation acknowledgements section. If you’re interested, you can download a copy here .

Psst... there’s more!
This post was based on one of our popular Research Bootcamps . If you're working on a research project, you'll definitely want to check this out ...
You Might Also Like:

Submit a Comment Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- Print Friendly
- Resources Home 🏠
- Try SciSpace Copilot
- Search research papers
- Add Copilot Extension
- Try AI Detector
- Try Paraphraser
- Try Citation Generator
- April Papers
- June Papers
- July Papers

How to Write Acknowledgement in Research Paper

Table of Contents
Writing an acknowledgement in a research paper is an integral part of the process. It is a formal way of expressing gratitude to the individuals and institutions that contributed to the completion of your research.
This section, though not mandatory, holds significant value as it acknowledges the efforts of those who assisted you in the successful completion of your project. In this comprehensive guide, we will delve into the intricacies of writing an effective acknowledgement for your research paper.
Introduction
Acknowledgements serve a crucial role in research papers . They not only express gratitude but also provide a sense of credibility to your work. Acknowledging the contributions of others shows that your research is a collective effort, which can enhance the perceived validity of your findings.
Moreover, acknowledgements can also serve as a platform for you to demonstrate your professional courtesy and respect for the individuals and institutions that have supported your research. This can help in fostering positive relationships, which can be beneficial for your future research endeavors.
Who to acknowledge in your research paper ?
Deciding who to acknowledge in your research paper can be a challenging task. It is important to ensure that you do not overlook anyone who has contributed to your research. Here are some categories of individuals and institutions that you might consider acknowledging:
Academic advisors and supervisors:
Your academic advisors and supervisors are likely to be your first point of contact for guidance and support during your research. They provide valuable insights, feedback, and direction, which can significantly influence the outcome of your research.
Therefore, acknowledging them in your research paper is a way of expressing your gratitude for their assistance and guidance. It also shows your respect for their expertise and dedication to your research.
Research participants and collaborators
Research participants and collaborators play a crucial role in the success of your research. They provide the data or information necessary for your research, making their contribution invaluable.
Acknowledging them in your research paper is a way of showing your appreciation for their time and effort. It also symbolizes your respect for their contribution to your research.
Funding bodies and institutions:
Funding bodies and institutions provide the financial support necessary for conducting your research. Without their support, it might be challenging to carry out your research effectively.
Therefore, acknowledging them in your research paper is a way of expressing your gratitude for their financial support. It also shows your appreciation for their trust in your research capabilities.
How to write acknowledgements for your research paper?
Writing acknowledgements for your research paper involves more than simply listing names. It requires a thoughtful and sincere expression of gratitude. Here are some steps to guide you in writing effective acknowledgements:
Start with the most significant contributions:
Begin your acknowledgements by expressing gratitude to those who have made the most significant contributions to your research. This could be your academic advisors, supervisors, or funding bodies. Starting with the most significant contributions helps to set the tone for the rest of your acknowledgements.
Ensure that you express your gratitude sincerely and professionally. Avoid using overly emotional or informal language as this can undermine the professionalism of your acknowledgements.
Acknowledge other contributors:
After acknowledging the most significant contributors, proceed to acknowledge other individuals and institutions that have supported your research. This could include research participants, collaborators, and other supportive individuals or institutions.
When acknowledging these contributors, be sure to express your gratitude sincerely and professionally. Also, ensure that you acknowledge each contributor individually to show your appreciation for their unique contribution.
Use appropriate language and tone:
The language and tone you use in your acknowledgements can significantly influence how they are perceived. Therefore, it is important to use appropriate language and maintain a professional tone throughout your acknowledgements.
Use formal language and avoid using jargon or colloquial expressions. Also, maintain a consistent tone throughout your acknowledgements to ensure that they are coherent and easy to read.
Examples of acknowledgements in research papers
Here are a few examples that demonstrate how to acknowledge different contributors effectively:
"I would like to express my deepest gratitude to my advisor, Professor ABC, for his invaluable guidance and support throughout this research. His expertise and dedication have been a source of inspiration and motivation."
Research participants and collaborators:
"I am deeply grateful to all the participants who generously shared their time and experiences for this research. Their contributions have been instrumental in the success of this study."
"This research was made possible by the generous funding from ABC Foundation. I am profoundly grateful for their support and trust in my research capabilities."
Writing acknowledgements in a research paper is a thoughtful process that requires careful consideration of who to acknowledge and how to express gratitude. By following the guidelines and examples provided in this article, you can write effective acknowledgements that reflect your appreciation and respect for the contributions of others to your research.
Remember, acknowledgements are more than just a formality. They are an opportunity to express your gratitude and respect for the individuals and institutions that have supported your research journey. So, take the time to write acknowledgements that are sincere, professional, and reflective of your gratitude.
Frequently Asked Questions
In a research paper, the acknowledgment section is where the author shows appreciation to those who helped with the research. It's usually found at the start of the paper, before the main text begins. However, the exact location varies depending on the university guidelines.
Citations are formal ways to acknowledge the sources you used, while acknowledgments are more personal and can serve as a confidential way to thank someone for their help or contribution.
The purpose of including acknowledgement is to express gratitude to everyone who assisted with the research but didn't meet the criteria for being listed as an author.
There is no minimum length for writing an acknowledgement in a research paper but it should not be more than one page.
If you conducted the research entirely by yourself and received no assistance or support from others, it's not necessary to include acknowledgements. However, if you received any form of support or assistance, even if minimal, it's appropriate to acknowledge it.
Yes, it's appropriate to acknowledge funding sources in your research.
You might also like

Boosting Citations: A Comparative Analysis of Graphical Abstract vs. Video Abstract

The Impact of Visual Abstracts on Boosting Citations

Introducing SciSpace’s Citation Booster To Increase Research Visibility
- SpringerLink shop
Acknowledgments and References
Acknowledgments.
This usually follows the Discussion and Conclusions sections. Its purpose is to thank all of the people who helped with the research but did not qualify for authorship (check the target journal’s Instructions for Authors for authorship guidelines). Acknowledge anyone who provided intellectual assistance, technical help (including with writing and editing), or special equipment or materials.
TIP: The International Committee of Medical Journal Editors has detailed guidelines on who to list as an author and who to include in the Acknowledgments that are useful for scientists in all fields.
Some journals request that you use this section to provide information about funding by including specific grant numbers and titles. Check your target journal’s instruction for authors for specific instructions. If you need to include funding information, list the name(s) of the funding organization(s) in full, and identify which authors received funding for what.
As references have an important role in many parts of a manuscript, failure to sufficiently cite other work can reduce your chances of being published. Every statement of fact or description of previous findings requires a supporting reference.
TIP: Be sure to cite publications whose results disagree with yours. Not citing conflicting work will make readers wonder whether you are really familiar with the research literature. Citing conflicting work is also a chance to explain why you think your results are different.
It is also important to be concise. You need to meet all the above needs without overwhelming the reader with too many references—only the most relevant and recent articles need to be cited. There is no correct number of references for a manuscript, but be sure to check the journal’s guidelines to see whether it has limits on numbers of references.
TIP: Never cite a publication based on what you have read in a different publication (such as a review), or based only on the publication’s abstract. These may mislead you and readers. Read the publication itself before you cite it, and then check the accuracy of the citation again before submitting your manuscript.
You should reference other work to:
- Establish the origin of ideas
When you refer to an idea or theory, it is important to let your readers know which researcher(s) came up with the idea. By citing publications that have influenced your own work, you give credit to the authors and help others evaluate the importance of particular publications. Acknowledging others’ contributions is also an important ethical principle.
- Justify claims
In a scientific manuscript, all statements must be supported with evidence. This evidence can come from the results of the current research, common knowledge, or from previous publications. A citation after a claim makes it clear which previous study supports the claim.
- Provide a context for your work
By highlighting related works, citations help show how a manuscript fits into the bigger picture of scientific research. When readers understand what previous studies found and what puzzles or controversies your study relates to, they will better understand the meaning of your work.
- Show there is interest your field of research
Citations show that other researchers are performing work similar to your own. Having current citations will help journal editors see that there is a potential audience for your manuscript.
Back │ Next

- Acknowledgements for PhD Thesis and Dissertations – Explained
- Doing a PhD
The Purpose of Acknowledgements
The acknowledgement section of a thesis or dissertation is where you recognise and thank those who supported you during your PhD. This can be but is not limited to individuals, institutions or organisations.
Although your acknowledgements will not be used to evaluate your work, it is still an important section of your thesis. This is because it can have a positive (or negative for that matter) influence the perception of your reader before they even reach the main body of your work.
Who Should I Acknowledge?
Acknowledgements for a PhD thesis will typically fall into one of two categories – professional or personal.
Within these categories, who you thank will ultimately be your decision. However, it’s imperative that you pay special attention to the ‘professional’ group. This is because not thanking someone who has played an important role in your studies, whether it be intentional or accidental, will more often than not be seen as a dismissal of their efforts. Not only would this be unfair if they genuinely helped you, but from a certain political aspect, it could also jeopardise any opportunities for future collaborations .
Professional Acknowledgements
This may include, but is not limited to:
- Funding bodies/sponsorship providers
- Supervisors
- Research group and lab assistants
- Research participants
- Proofreaders
Personal Acknowledgements
- Key family members and friends
- Individuals who inspired you or directly influenced your academic journey
- Anyone else who has provided personal support that you would like to mention
It should be noted that certain universities have policies which state only those who have directly supported your work, such as supervisors and professors, should be included in your acknowledgements. Therefore, we strongly recommend that you read your university guidelines before writing this section of your thesis.
How to Write Acknowledgements for PhD Thesis
When producing this section, your writing style can be more informal compared to the rest of your thesis. This includes writing in first person and using more emotive language. Although in most cases you will have complete freedom in how you write this section of your thesis, it is still highly advisable to keep it professional. As mentioned earlier, this is largely because it will be one of the first things your assessors will read, and so it will help set the tone for the rest of your work.
In terms of its structure, acknowledgements are expected to be ordered in a manner that first recognises the most formal support before moving onto the less formal support. In most cases, this follows the same order that we have outlined in the ‘Who Should I Thank’ section.
When thanking professionals, always write out their full name and provide their title. This is because although you may be on a first-name basis with them, those who read your thesis will not. By providing full names and titles, not only do you help ensure clarity, but it could also indirectly contribute to the credibility of your thesis should the individual you’re thanking be well known within your field.
If you intend to include a list of people from one institution or organisation, it is best to list their names in alphabetical order. The exception to this is when a particular individual has been of significant assistance; here, it would be advisable to list them.
How Long Should My Acknowledgements Be?
Acknowledgements vary considerably in length. Some are a single paragraph whilst some continue for up to three pages. The length of your acknowledgement page will mostly depend on the number of individuals you want to recognise.
As a general rule, try to keep your acknowledgements section to a single page. Although there are no word limits, creating a lengthy acknowledgements section dilutes the gratitude you’re trying to express, especially to those who have supported you the most.
Where Should My Acknowledgements Go?
In the vast majority of cases, your acknowledgements should appear directly after your abstract and before your table of contents.
However, we highly advise you to check your university guidelines as a few universities set out their own specific order which they will expect you to follow.
Phrases to Help You Get Started

We appreciate how difficult it can be to truly show how grateful you are to those who have supported you over the years, especially in words.
To help you get started, we’ve provided you with a few examples of sentences that you can complete or draw ideas from.
- I am deeply grateful to XXX…
- I would like to express my sincere gratitude to XXX…
- I would like to offer my special thanks to XXX…
- I would like to extend my sincere thanks to XXX…
- …for their assistance at every stage of the research project.
- …for their insightful comments and suggestions.
- …for their contribution to XXX.
- …for their unwavering support and belief in me.
Thesis Acknowledgement Examples
Below are three PhD thesis acknowledgment samples from which you can draw inspiration. It should be noted that the following have been extracted from theses which are freely available in the public domain. Irrespective of this, references to any individual, department or university have been removed for the sake of privacy.
First and foremost I am extremely grateful to my supervisors, Prof. XXX and Dr. XXX for their invaluable advice, continuous support, and patience during my PhD study. Their immense knowledge and plentiful experience have encouraged me in all the time of my academic research and daily life. I would also like to thank Dr. XXX and Dr. XXX for their technical support on my study. I would like to thank all the members in the XXX. It is their kind help and support that have made my study and life in the UK a wonderful time. Finally, I would like to express my gratitude to my parents, my wife and my children. Without their tremendous understanding and encouragement in the past few years, it would be impossible for me to complete my study.
I would like to thank my supervisors Dr. XXX and Dr. XXX for all their help and advice with this PhD. I would also like to thank my sisters, whom without this would have not been possible. I also appreciate all the support I received from the rest of my family. Lastly, I would like to thank the XXX for the studentship that allowed me to conduct this thesis.
I would like to thank my esteemed supervisor – Dr. XXX for his invaluable supervision, support and tutelage during the course of my PhD degree. My gratitude extends to the Faculty of XXX for the funding opportunity to undertake my studies at the Department of XXX, University of XXX. Additionally, I would like to express gratitude to Dr. XXX for her treasured support which was really influential in shaping my experiment methods and critiquing my results. I also thank Dr. XXX, Dr. XXX, Dr. XXX for their mentorship. I would like to thank my friends, lab mates, colleagues and research team – XXX, XXX, XXX, XXX for a cherished time spent together in the lab, and in social settings. My appreciation also goes out to my family and friends for their encouragement and support all through my studies.
Browse PhDs Now
Join thousands of students.
Join thousands of other students and stay up to date with the latest PhD programmes, funding opportunities and advice.

The Savvy Scientist
Experiences of a London PhD student and beyond
Thesis acknowledgements: Samples and how to write your own thesis or dissertation acknowledgements

Writing a thesis can be tricky. That’s why I’m starting a new series covering each section of the thesis, from thesis acknowledgements all the way to conclusions. I’ll be guiding you through the whole process, from what to include in your thesis to how to write it, along with examples from defended theses to help you to write your own.
We’ll begin by covering thesis acknowledgments. The acknowledgements section appears at the start of the thesis so it is often one of the first parts that everyone tries to tackle. As this will likely be your first taste of your thesis it can often feel quite intimidating to write!
Thankfully it’s also one of the easiest parts of the thesis to complete, which may help to give you a boost for the rest.
In this post we’ll cover everything to do with thesis acknowledgements: samples, what to include and how to write them. At the end I’ll also outline a 60 minute exercise which will get you preparing a first draft of your own! I’ve also got a similar post to craft your thesis title, which you can check out here .
I’m writing this post with a PhD thesis in mind but it could work just as well if you’re looking for help including acknowledgements in your Master’s or undergraduate thesis/ dissertation.
What is the purpose of the acknowledgements section in a thesis?
The acknowledgements section of your thesis is an opportunity to reflect on the people who have supported and shaped your PhD experience.
Don’t worry, although your examiners will be interested to read your acknowledgements section, you won’t really get judged on it in your PhD viva. This section is for you to share as little, or as much, as you want about everyone involved in your PhD journey.
The acknowledgements are a very personal section of your thesis and each PhD student will have different things they want to include. For example, many people wonder: How do I thank my family in a thesis? And the acknowledgements section is the answer!
Note – You can also use a thesis dedication to thank your family. This is a separate section to your thesis acknowledgements and is entirely optional. It’s usually just a single line, just like you might find at the front of some books. Most people don’t include a separate dedication section but you can if you want to go that extra step.
What to include in your thesis acknowledgements
There are usually no formal requirements dictating what to include in your acknowledgements. However, do double check for any potential rules at your specific institution.
In general the acknowledgements are the section of your thesis where you have some creative liberty and are not bound by rigid research protocols or guidelines.
Many students choose to use the acknowledgements section to thank people (or organisations) who:
- Introduced them to the topic
- Helped with their PhD application
- Funded the project
- Supervisors
- Technicians
- Partners, friends or family
- Or anyone else who made an impression along the way!
But remember, you can include whatever you want! For example in my own PhD acknowledgements, which you’ll read further down this post, I thanked the university for providing a green outdoor space for us.
Acknowledge whoever and whatever influenced your own PhD experience.
You may find it helpful to start by writing a list of everyone you wish to thank.
How do you write an acknowledgements section?
Since there are no guidelines to worry about, it is really up to you how you write your own thesis acknowledgements. You have a lot of freedom for what to include and how to write it.
However you may find the following suggested phases helpful as a starting point.
Who you want to thank…
- “First and foremost, I would like to express my sincere gratitude to…”
- “I must thank…”
- “A special thanks to…”
- “I would like to highlight two truly exceptional people from…”
- “I want to thank…”
- “In addition, I would like to mention”
- “I would also like to extend my thanks to…”
- “I want to give my deepest appreciation to…”
- “Finally, but the most importantly, I would like to thank…”
…then, why you want to thank them
It can be nice to also include why you’re thanking these people, using phrases such as:
- “…for the opportunity to be a part of this project”
- “…for always being there when I needed his support, reviewing my progress constantly, and guiding me through my PhD studies”
- “….for being a great bunch of people in and out of the lab”
- …”for all the guidance, support and outstanding feedback”
- “… who took their time to help teach me…”
- “…for her unlimited support and unconditional guidance during my PhD journey”
- “…were always there for discussions about anything that I was unsure on”
- “…whom has offered invaluable advice that will benefit me throughout my life”
- “…for supporting me since my undergraduate, and for the valuable discussions we had along the road”
- “…for making the past 4 years much more enjoyable and keeping me sane throughout the whole process”
Here is a whole example from an accepted PhD thesis:
Firstly, I want to thank [supervisor’s name(s)] for giving me the opportunity to work on this project, providing valuable guidance and feedback, and challenging me to grow as a scientist. Excerpt from Dr Wane’s thesis acknowledgements, available via this page or use this direct download link .
Some people will choose to use full names and titles for any professional acknowledgements and first names for any personal ones. Again, this is up to you.
To help illustrate the variety of thesis acknowledgement formats, we’ll shortly be coming on to some examples of acknowledgment sections from successfully defended theses.
Before then I want to cover some of the main questions relating to how to write your own thesis acknowledgements section:
How long should you spend writing your thesis acknowledgements?
My suggestion is to spend only an hour or two making a first draft. I suggest doing this well ahead of your final deadline so that you have time to come back to it. Even so, I’d certainly look to spend far less than one day’s work on it in total.
It is a “nice to have” and means a lot to a lot of people, but remember you’re really only writing this section for yourself. I probably spent about two hours writing mine in total, simply because it wasn’t a priority for me.
What order should you write your acknowledgements in?
A typical way to write your acknowledgements is to go from the most formal/academic relationships to the least.
It is normal to start with any funding bodies, then formal people like your PhD supervisors, then move through labmates, friends and family. But again, there are generally no rules!
How long should the acknowledgements section be?
You can include as much or as little as you want. My own PhD acknowledgements section was just under a page long and it consisted of 386 words or 1892 characters (without spaces).
Here is how it was formatted:

But let’s not just look at my thesis. Using Imperial’s publicly accessible database I went through 25 published PhD theses for you.
The average (mean) length of these 25 theses was 365 words and 1793 characters without spaces. Writing an acknowledgements section of length 350-450 words was the most common:

The shortest acknowledgements sections was 122 words(653 characters) long. The longest one consisted of 1022 words and 5082 characters. Hopefully this illustrates that you’re not really bound by any limits. Write as much or as little as you want for this section.
Sample thesis acknowledgements
My own phd thesis acknowledgement.
My own PhD thesis is available here *, the acknowledgements section is on page 5. Here is the complete version of my acknowledgements section:
I would like to acknowledge both EPSRC and the Class of 1964 Scholarship for their financial support. It has been an honour to be the inaugural recipient of the Class of 1964 Scholarship and I am indebted to the donors in providing me complete academic freedom in this research. An immense thank you to my PhD supervisors: Jonathan Jeffers, Ulrich Hansen and Julian Jones. Support and guidance throughout the project from you all has been invaluable. JJ in particular you’ve been a fantastic primary supervisor. Thank you to all the academics who helped me get to this stage. The late Dr Kajal Mallick and his Biomedical Materials course at the University of Warwick was a huge influence and without which I would have never followed this path. My “pre-doc” supervisors in Dr Helen Lee of University of Cambridge and in particular the remarkable Prof Judith Hall OBE of Cardiff University from whom I learned so much. Thanks to Alison Paul and Michael Lim for being so supportive when I was considering applying for PhDs. It has been an amazing experience working between two research groups across different departments, thanks to everyone from the Biomechanics and JRJ groups I’ve worked with and from whom I’ve learned so much. Thank you of course to the Hybrids team I’ve worked so closely on this project with: Fra, Gloria, Agathe, Maria, Silvia, it’s been great fun working with you all! Gloria in particular thanks for you all your help, support and friendship: your inclusivity is appreciated by many. Saman, I’ve been so pleased to have you working on DVC with me and being able to discuss ideas with you really has been invaluable. I am grateful to everyone I’ve collaborated with externally: Farah, Amin and Brett (Natural History Museum) plus Andy and Behzad (Royal Veterinary College), thank you all for your support and input. Thanks also to everyone I’ve met through the Environmental Society at Imperial in particular Chelcie: your friendship and support have added a lot to my life. Thanks to Imperial for providing space for the ESoc garden, taking a break and enjoy nature in this space has certainly improved my work. Thanks of course to my family for their support. Finally, thank you Jo for always being so supportive and helping me every step of the way. My PhD thesis, available here . Acknowledgements are on page 5.
*For me the thesis was a means to an end. I wanted my PhD and didn’t want to spend too long agonising over each page. Therefore, it is possible there are typos in there, if you read any of it: firstly well done, I haven’t looked at it much since submitting the final copy, secondly, please don’t tell me about any typos you find!
Other PhD thesis acknowledgement examples
Below are the other 24 published and openly accessible STEM PhD theses I found for this article.
For each person’s thesis, either follow the first link to be taken to the landing page or follow the second link to directly download their thesis: I gave you a choice in case you don’t want stuff to start downloading automatically from a random text link!

The list is formatted as follows:
- [Link to thesis page on repository], [which page the acknowledgements appear on], [direct link to download the thesis]
- Dr Shipman’s thesis , for the acknowledgements go to page 3. Direct download here .
- Longest acknowledgements section of the list at 1022 words.
- Dr Li’s thesis , page 11. Direct download here .
- Dr Podgurschi’s thesis , page 5. Direct download here .
- Dr Medjeral-Thomas’ thesis page 3. Direct download here .
- Dr Sztuc’s thesis , page 5. Direct download here .
- Dr Yap’s thesis , page 5. Direct download here .
- Dr Sukkar’s thesis , page 9. Direct download here .
- Dr Lo’s thesis , page 11. Direct download here .
- Dr Sullivan’s thesis , page 5. Direct download here .
- Dr Tawy’s thesis , page 3. Direct download here .
- Dr Wane’s thesis , page 2. Direct download here .
- Dr Addison’s thesis , page 4. Direct download here .
- Dr Wang’s thesis , page 5. Direct download here .
- Dr Sebest’s thesis , page 3. Direct download here .
- Dr Hopkins’ thesis , page 7. Direct download here .
- Dr Bates’s thesis , page 4. Direct download here .
- Dr Somuyiwa’s thesis , page 6. Direct download here .
- Dr Reynolds’ thesis , page 5. Direct download here .
- My labmate’s thesis, who wrote the acknowledgements in a different style to the rest by using bullet points.
- Shortest acknowledgements section of the list at 122 words.
- Dr Manca’s thesis , acknowledgements on page 5. Direct download here .
- Dr Liu’s thesis , page 5. Direct download here .
- Dr Hotinli’s thesis , page 7. Direct download here .
My top tips for writing your own thesis acknowledgements
- Don’t spend too long on them. The acknowledgements section is really not worth spending too much time on. Even worse, since they appear at the start of your thesis, it is tempting to write your acknowledgements first. This can be fine, or, it can be an opportunity for lots of unnecessary procrastination. Which I why I instead suggest that you…
- Write your acknowledgements at the end of your first draft of the thesis. There is no need to write your thesis in the order it is presented. If you write your acknowledgements at the end you’ll be less likely to spend precious time on a section which really doesn’t warrant too much brain power.
- Don’t stress about it. The acknowledgements are merely for yourself and for anyone close to you that you want to thank. There are far more important sections for you to be particular about!
- Remember: You can make changes after you submit the copy for your viva. As with everything in your thesis, you can make changes after you submit the thesis for your viva. The real “final” copy is when you submit your thesis to the university for archiving. Which is even more reason to not spend too much time writing it the first time around.
Draft your own thesis or dissertation acknowledgements in 60 minutes
Hopefully you now feel inspired to start writing your own thesis acknowledgments!
For the exercise below I’d suggest setting a stop-watch on your phone and move on to the next section when the alarm goes, even if you’ve not fully finished. The aim is to have a rough draft at the end which you can polish off at a later point in time.
- Read a few of the example thesis acknowledgements above to get a feel for the structure ( 15 mins )
- List everyone (or everything!) you wish to thank – including any personal and professional acknowledgements in addition to funding bodies if relevant ( 10 mins )
- Decide on a rough order in which to thank them ( 5 mins )
- Craft some sentences using the phrases mentioned above ( 30 mins )
Congratulations you’re now well on your way to having one section of your PhD thesis completed!
I hope this post has been useful for constructing your own thesis or dissertation acknowledgements. It is the first in a series of posts aiming to help your thesis writing by delving into each section in depth. Be sure to let me know if you have any questions or suggestions for other content which you would find useful.
Subscribe below to stay updated about future posts in the series:
Share this:
- Click to share on Facebook (Opens in new window)
- Click to share on LinkedIn (Opens in new window)
- Click to share on Twitter (Opens in new window)
- Click to share on Reddit (Opens in new window)
Related Posts

Minor Corrections: How To Make Them and Succeed With Your PhD Thesis
2nd June 2024 2nd June 2024

How to Master Data Management in Research
25th April 2024 27th April 2024

Thesis Title: Examples and Suggestions from a PhD Grad
23rd February 2024 23rd February 2024
Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Notify me of follow-up comments by email.
This site uses Akismet to reduce spam. Learn how your comment data is processed .
Privacy Overview
- Link to facebook
- Link to linkedin
- Link to twitter
- Link to youtube
- Writing Tips
How to Write Acknowledgements
- 2-minute read
- 25th March 2015
If you are near the end of your thesis, you can start to think about putting on the finishing touches . One thing you will have to do here is write your acknowledgements. A lot of people worry about this, but there’s really no need. As we are about to show, it’s actually quite simple!
What Are Acknowledgements?
The acknowledgement section of a thesis is where you can thank everyone who has helped you in your research. It is typically located at the beginning of your thesis, right after the contents page, and shouldn’t really be more than one or two pages long. The best thing to do is to keep it concise.
Who Should I Thank?
Think about the people who were of crucial importance during your research. This could include friends, family or professors, or even volunteers who have taken part in your research. It’s also important to acknowledge professional bodies who have given you funding or other help.
How Should I Write It?
There are no strict requirements for the tone of your acknowledgements; it’s essentially the one section where you can be a little bit more informal! Try to make sure the tone fits the person or organization you’re thanking, though, maintaining a formal approach when addressing funding bodies or other official groups.
Find this useful?
Subscribe to our newsletter and get writing tips from our editors straight to your inbox.
Other than trying to be concise, the main thing you should keep in mind is varying your language, if only because starting every sentence with “I would like to thank…” will sound a bit dull.
There are several ways you could word your thanks. A few suggestions are:
- This research would not have been possible without…
- My sincere thanks go to…
- I am grateful to…
- Heartfelt thanks to…
- I would like to express my gratitude to…
- Appreciation is due to…
- I acknowledge the contribution of…
- I am indebted to…
And that’s all there is to it! Relax and enjoy writing your acknowledgements. If you’ve got this far in the thesis-writing process, you’re over the toughest bit now! For more information about writing a dissertation or thesis, read our full dissertation writing guide .
Share this article:
Post A New Comment
Got content that needs a quick turnaround? Let us polish your work. Explore our editorial business services.
9-minute read
How to Use Infographics to Boost Your Presentation
Is your content getting noticed? Capturing and maintaining an audience’s attention is a challenge when...
8-minute read
Why Interactive PDFs Are Better for Engagement
Are you looking to enhance engagement and captivate your audience through your professional documents? Interactive...
7-minute read
Seven Key Strategies for Voice Search Optimization
Voice search optimization is rapidly shaping the digital landscape, requiring content professionals to adapt their...
4-minute read
Five Creative Ways to Showcase Your Digital Portfolio
Are you a creative freelancer looking to make a lasting impression on potential clients or...
How to Ace Slack Messaging for Contractors and Freelancers
Effective professional communication is an important skill for contractors and freelancers navigating remote work environments....
3-minute read
How to Insert a Text Box in a Google Doc
Google Docs is a powerful collaborative tool, and mastering its features can significantly enhance your...

Make sure your writing is the best it can be with our expert English proofreading and editing.
Reference management. Clean and simple.
Dissertation acknowledgments [with examples]

What are dissertation acknowledgements?
What to consider when writing your dissertation acknowledgments, who to thank in your dissertation acknowledgments, what (and what not) to write in your dissertation acknowledgments, good examples of dissertation acknowledgments, a final word on writing dissertation acknowledgments: have fun, frequently asked questions about dissertation acknowledgments, related articles.
While you may be the sole author of your dissertation, there are lots of people who help you through the process—from your formal dissertation advisors to the friends who may have cooked meals so that you could finish your last chapter . Dissertation acknowledgments are a chance to thank everyone who had a hand in the completion of your project.
Dissertation acknowledgments are a brief statement of your gratitude to advisors, professors, peers, family, and friends for their help and expertise.
In this guide, we’ll cover:
- the most important things to consider when you’re writing your dissertation acknowledgments
- who to thank in your dissertation acknowledgments
- what (and what not) to write in your dissertation acknowledgments
- short examples of dissertation acknowledgments
Once you’re at the stage where you’re writing your dissertation acknowledgments, you may be tempted to kick back and relax. After all, the hard part of writing the dissertation itself is over and a list of thanks should be simple to churn out.
However, the acknowledgments are an important part of your overall work and are something that most people who read your dissertation, including prospective employers, will look at.
Tip: The best dissertation acknowledgements are concise, sincere, and memorable.
Approach this part of the process, brief as it may be compared to the long haul of writing the dissertation, with the same high level of care and attention to detail. It’s an explicit and permanent statement of who made a real impact on your work and contributed to your academic success.
Plus, the people you thank are often deeply moved by being included—some even go so far as to frame the acknowledgments. Aim to make yours sincere, memorable and something that people will be touched by.
First things first: who should you include in your dissertation acknowledgments? If you’re not sure who to thank, try the brainstorming technique to generate some ideas. Consider these two approaches:
- Make a list of everyone, both professional and personal, who was involved at any point during your work on your dissertation, and then thin down the list from there.
- Make a list of the pivotal aspects of your process and think about who was involved and how they helped.
As you select the people and groups to include in your dissertation acknowledgments, keep in mind that it’s essential to acknowledge your supervisor and anyone else with a visible connection to your work.
It’s an unfortunate reality that not every supervisor goes above and beyond to provide feedback and guidance to the students they are supposed to supervise. However, leaving them out, even if you personally felt disappointed by their involvement or lack thereof, could be seen as a snub.
You should end up with a fairly short list of people to thank. While being mindful of professional etiquette and personal feelings, be choosy about who makes the final cut since your acknowledgments should be limited to no more than a page.
Now that you have your list of people and groups to thank, it’s time to start writing. Before your first pen or keystroke, however, check your university’s guidelines as your institution may have specific rules around what can and cannot be included.
The standard practice is to begin with the formal and then progress to the informal, so the first people to mention would be:
- supervisors
- committee members
- other professional contacts
Use their full names and titles and go into brief detail about how they contributed to your work.
Once those are done, you can move on to the personal thanks, which can include friends, family, even pets. If you are so inclined, it is also considered appropriate to thank God or make mention of spiritual support.
You may also choose to inject a little humor at this point, but don’t get carried away and definitely don’t include sarcasm or critical comments of any kind, including self-critical ones. Remember that the acknowledgments precede your dissertation, so you want to be taken seriously.
A couple more basics that are essential when creating your acknowledgments:
- Position: Acknowledgments should be placed after the title page and before the abstract.
- Perspective: Write from the first-person perspective and speak in your own voice.
A really good way to get a sense of how to write your own dissertation acknowledgments is to read ones written by others. Notice which ones you respond particularly well to and use them as a model upon which to base your own.
Here are some good examples to help you get started:
I couldn’t have reached this goal without the help of many people in my life. I’d like to take this opportunity to thank them for their support.
First, my sincere thanks to my dissertation committee. The value of their guidance cannot be overstated. Dr. Elaine Gooding and Dr. Matthew Hunter provided much wisdom that helped me chart my course. I couldn’t have asked for a better supervisor than Dr. Fiona Moore, whose knowledge and experience guided me every step of the way.
Next, I’d like to thank my partner, Elliott. Your votes of confidence kept me going when my spirits dipped. I couldn’t have done this without you.
Last but not least, I’d like to acknowledge the emotional support provided by my family and friends. We made it to the top of the mountain! I look forward to celebrating with all of you.
This example is shorter, but still contains the key components:
Several people played a decisive role in my success and I would like to take this opportunity to thank them.
My chair, Dr. Ronald Saulk, provided invaluable support and infinite patience and I am truly grateful for all of his wisdom and guidance. I also owe the entire staff of the Wilhelm Library a debt of gratitude. From tracking down books and arranging for interlibrary loans to keeping the coffee maker in the lobby well-stocked and in good working order, they offered the practical help and kind gestures that made all the difference.
I’d also like to thank my family and God, for always being there for me.
One final piece of advice: enjoy this process. Writing a dissertation doesn’t happen every day, and the opportunity to acknowledge the important people in your life in a published format is as rare as it is wonderful.
What’s more, this part of your dissertation is unlike any other. It’s unbounded by the conventions that apply to the formal work. It’s a chance to really flex some creative muscle and let your personality shine through. So make the most of it and have fun!
In your dissertation acknowledgments, you thank everyone who has contributed to your work or supported you along the way. Who you want to thank is a very personal choice, but you should include your supervisors and anyone else with a visible connection to your work. You may also thank friends, family, and partners.
First, you need to come up with a list of people you want to thank in your dissertation acknowledgments. As a next step, begin with the formal and then progress to the informal, so the first people to mention would be supervisors, mentors, committees, and other professional contacts. Then, you can move on to the personal thanks, which can include friends, family, even pets.
Who you acknowledge in your dissertation is ultimately up to you. You should, however, thank your supervisor and anyone else with a visible connection to your work. Leaving them out, even if you personally felt disappointed by their involvement or lack thereof, could be seen as a snub. In addition, you can thank friends, partners or family.
There are many ways so you can acknowledge your dissertation supervisor. Some examples can be found in this article above. If you need more examples, you can find them here .
While acknowledgments are usually more present in academic theses, they can also be a part of research papers. In academic theses, acknowledgments are usually found at the beginning, somewhere between abstract and introduction. In research papers, acknowledgments are usually found at the end of the paper.


How to Draft the Acknowledgment Section of a Manuscript
What is the Purpose of the Acknowledgements Section in a Research Paper?
The acknowledgment section is an integral part of all academic research papers. It provides appropriate recognition to all contributors for their hard work. We discuss here, the relevant guidelines for acknowledging contributors.
Defining Who Is Acknowledged
The acknowledgment section helps identify the contributors responsible for specific parts of the project. It can include:
- Non-authors (colleagues, friends, supervisor, etc.)
- Funding sources
- Editing services ,
- Administrative staff
In academic writing, the information presented in the acknowledgment section should be kept brief. It should only mention people directly involved with the project. In other words, one should not consider thanking ones’ parents for moral and financial support.
Acknowledging contributors is necessary. However, you must know the difference between an author and a contributor . The International Committee of Medical Journal Editors ( ICMJE ) defines four criteria to assign authorship.
He or she has to have
- Made substantial conceptual or design contributions or gathered and analyzed important data, and
- Either helped draft or critically revise the paper in keeping with important intellectual content, and
- Provided final approval before publishing, and
- Agreed to be accountable for the accuracy of the work
These authors and their affiliations will be listed at the beginning of the paper. The “corresponding author” will also be listed a second time and will directly correspond with the journal to ensure documentation requirements are met.
Many journals now ask that you provide the role of each author in your acknowledgment section. For example, a typical statement of authors’ contributions might be as follows (note that only last names are used unless ambiguous):
Smith conducted the data analysis and created the tables and figures. Jones provided his technological expertise for GIS tracking. Johnson provided a factual review and helped edit the manuscript.
This type of acknowledgment provides your reader with a good sense of who was responsible for each part of your research and manuscript.

Non-Author Contributors
There are many people involved in a research project who are not authors but have provided valuable contributions. For example, one person’s responsibility might be to seek project funding; another’s might be to supervise laboratory staff. A few others might have provided valuable services such as technical editing and writing or offering help in reviewing and revising the manuscript for grammar and syntax. These people should also be mentioned in the acknowledgment section of your manuscript.
Acknowledgment should also be provided for writing assistance, technical editing, language editing, and proofreading . Therefore, editing companies need to be duly acknowledged in professionally edited manuscripts as per the ICMJE guidelines.
It is necessary to acknowledge editing companies in professionally edited manuscripts, even though these companies are paid for their work.
Acknowledgment Format
Unlike the main body of your paper, the format for your acknowledgment section can be more personal. It is permissible to use personal pronouns in this section. For example,
I thank the following individuals for their expertise and assistance throughout all aspects of our study and for their help in writing the manuscript.
Keep in mind that many guidelines indicate that funding sources be listed separately from the acknowledgment section. In addition, the sources (funding agencies) might have specific guidelines that you must follow. Please be sure to comply with these sources and your author guidelines.
For more information on authors and contributors , read articles on the Enago Academy website.
What types and formats of acknowledgments have you incorporated into your manuscripts? Please share your thoughts in the comments section below. Do you need help with manuscript editing ? Make sure you visit enago.com today!
Thanks!!! This information helped me a lot in finishing my research paper.
This information was very useful for preparing this paper.
We would like to thank Enago (www.enago.com) for the English language review.”
Rate this article Cancel Reply
Your email address will not be published.

Enago Academy's Most Popular Articles

- AI in Academia
- Infographic
- Manuscripts & Grants
- Reporting Research
- Trending Now
Can AI Tools Prepare a Research Manuscript From Scratch? — A comprehensive guide
As technology continues to advance, the question of whether artificial intelligence (AI) tools can prepare…

Abstract Vs. Introduction — Do you know the difference?
Ross wants to publish his research. Feeling positive about his research outcomes, he begins to…

- Old Webinars
- Webinar Mobile App
Demystifying Research Methodology With Field Experts
Choosing research methodology Research design and methodology Evidence-based research approach How RAxter can assist researchers

- Manuscript Preparation
- Publishing Research
How to Choose Best Research Methodology for Your Study
Successful research conduction requires proper planning and execution. While there are multiple reasons and aspects…

Top 5 Key Differences Between Methods and Methodology
While burning the midnight oil during literature review, most researchers do not realize that the…
Discussion Vs. Conclusion: Know the Difference Before Drafting Manuscripts
Annex Vs. Appendix: Do You Know the Difference?

Sign-up to read more
Subscribe for free to get unrestricted access to all our resources on research writing and academic publishing including:
- 2000+ blog articles
- 50+ Webinars
- 10+ Expert podcasts
- 50+ Infographics
- 10+ Checklists
- Research Guides
We hate spam too. We promise to protect your privacy and never spam you.
I am looking for Editing/ Proofreading services for my manuscript Tentative date of next journal submission:

What would be most effective in reducing research misconduct?
- How it works

How to Write Dissertation Acknowledgements
Published by Owen Ingram at August 13th, 2021 , Revised On August 29, 2023
Acknowledging someone means thanking them. The acknowledgement section in a dissertation is used to express gratitude towards all those who have helped you prepare the dissertation.
Both professional and personal acknowledgements can be included in it. The acknowledgement section comes in between the title page and the abstract page . It is best suited to be on one page.
The writer has the liberty to use personal pronouns (I, we, my, etc) in this section, as this section allows more of an informal way of writing. Here is a quick guide to help you understand how to write acknowledgements for your own dissertation .
Dissertation Acknowledgement Example
Let’s start with an example, so you have an idea of the basics, to begin with.
How to Write Dissertation Acknowledgements?
“I am deeply grateful for the guidance and support of my supervisor, Dr. Smith, whose insights and feedback were invaluable throughout this journey. I also extend my appreciation to my family for their unwavering encouragement and to my friends for their understanding during this endeavor.”
“I am deeply grateful for the guidance and support of my supervisor, Dr. Smith, whose insights and feedback were invaluable throughout this journey. I also extend my appreciation to my family for their unwavering encouragement and to my friends for their understanding during this endeavour.”
You May Also Like
Your dissertation introduction chapter provides detailed information on the research problem, significance of research, and research aim & objectives.
Wish that you had more time to write your dissertation paper? Here are some practical tips for you to learn “How to get dissertation deadline extension”.
Dissertation conclusion is perhaps the most underrated part of a dissertation or thesis paper. Learn how to write a dissertation conclusion.
USEFUL LINKS
LEARNING RESOURCES

COMPANY DETAILS

- How It Works
Writing Acknowledgments for Your Research Paper
- Research Process
- Peer Review
In this article, we describe what types of contributions warrant mention in the acknowledgments section of a paper .
Updated on July 8, 2014

In another article , we discuss four criteria that must be met for an individual to qualify for manuscript authorship. In this article, we describe what types of contributions warrant mention in the acknowledgments section of a paper instead. The International Committee of Medical Journal Editors (ICMJE) describes several roles that merit acknowledgment, rather than authorship :
“acquisition of funding; general supervision of a research group or general administrative support; and writing assistance, technical editing, language editing, and proofreading.”
You should also acknowledge direct technical assistance, including help with animals, cells, equipment, patients, procedures, or techniques or provision of data, equipment, reagents, or samples, as well as more indirect assistance via intellectual discussions. Note that all of these contributions are typically more mechanical, indirect, and/or one-dimensional than those of authors. Additionally, some argue that individuals who provided help and could be chosen as a peer reviewer, leading to a potential conflict of interest, should be cited.
In any case, the ICMJE states that contributors may be cited individually or collectively and that their precise contributions should be specified.
e.g., “We thank Dr. X and Dr. Y for performing the surgeries” or “We thank the physicians who performed the surgeries"
Institutional affiliations may or may not be mentioned, depending on the journal's guidelines. Finally, the ICMJE encourages written permission from acknowledged individuals “because acknowledgment may imply endorsement.”
Funding sources should also be mentioned in the acknowledgments section, unless your target journal requires a separate section for this information. Whether the funding was partial or full, relevant grant numbers, and the author(s) who received the funding, if applicable, should be detailed as well. Note that acknowledging grants and fellowships is in fact required by many funding agencies and research institutions.
In contrast, contributions that are not specifically related to your research, including personal encouragement (e.g., by your friends or parents) and very general help (e.g., from a laboratory manager who purchases all supplies for your research group), should not be cited. Additionally, anonymous editors and peer reviewers are usually not thanked in the acknowledgments section; many journals (such as American Physical Society journals ) explicitly discourage this practice because it is difficult to comprehensively acknowledge all anonymous support and because this practice could potentially bias reviewers.
The writing style of acknowledgments sections may vary according to the journal, but generally, these sections are written in the first person and are as succinct as possible. A statement about conflicts of interest, citation of previous publication in poster or abstract form, and other information may also be included in this section, again depending on the journal. As you proceed through revisions for one journal or if you change your target journal, remember to reformat as necessary and to update your acknowledgments if additional help was obtained during the revision, such as with editing or new experiments.
Although an acknowledgments section may be appended to the end of your manuscript or relegated to a footnote, it is not a trivial component. By acknowledging all help received with your research, you are demonstrating your integrity as a researcher, which in turn encourages continued collaboration. You may also be bolstering your colleagues' careers, as being credited in an acknowledgments section is emerging as one of many gauges of a researcher's professional impact beyond citations (see ImpactStory , based on altmetrics ). Furthermore, information about who provided certain data, equipment, protocols, reagents, or samples may be of help to other researchers in your field.
This editing tip has hopefully elucidated what to include in the acknowledgments section of your manuscript and why this section is significant. If you have any comments or questions, please contact us . Best wishes in your research and writing!

Michaela Panter, PhD
See our "Privacy Policy"
- Translation
What to include in your Acknowledgments section
By charlesworth author services.
- Charlesworth Author Services
- 02 June, 2018
- Academic Writing Skills
Most academic papers have many people who have helped in some way in the preparation of the written version or the research itself. This could be someone from a sponsoring institution, a funding body, other researchers, or even family, friends or colleagues who have helped in the preparation. These people need to be mentioned in the Acknowledgments section of the paper.
Acknowledgments section in different academic documents
The Acknowledgments section is present in both a paper and an academic thesis . For papers, the Acknowledgments section is usually presented at the back, whereas in a thesis, this section is located towards the front of the manuscript and is commonly placed somewhere between the abstract and Introduction . However, the exact location varies between each university , as each establishment possesses its own style guide for theses and student submissions. So, it is always worthwhile consulting your university’s academic style guide before writing a manuscript for undergraduate/postgraduate submission.
Acknowledgments section in theses
For academic theses, there is no right or wrong way to acknowledge people, and who you want to acknowledge is down to personal preference. However, the common types of people authors acknowledge in their academic theses include:
- Supervisor’s contributions
- Research group (especially if the thesis in question is a master’s and the work is helped along by a PhD student)
- Support staff (laboratory technicians, etc.)
- Any students who undertook side projects with them (e.g. final year undergraduates, summer students, master’s students)
- Administrative staff (there can be a lot of bureaucracy for thesis submissions)
- Referees that got them onto the course (postgraduate only)
- Funding bodies
- Any collaboration with industry and the people they worked with at said establishment(s)
Acknowledgments section in journal papers
Now, whilst university manuscripts can include any combination of the above (including all and none in some cases), academic publications in journals more commonly acknowledge the same kind of people/organizations, but again it is up to the author(s) what they feel should be acknowledged; not every piece of help needs to be acknowledged, just the most useful/prevalent help. Also, acknowledgments should be written in the first person .
Examples of whom and what should be acknowledged in a journal publication are listed below:
- Direct technical help (e.g. supply of animal subjects, cells, equipment setup, methods , statistics/data manipulation, samples, chemicals/reagents, analytical/spectroscopy techniques)
- Indirect assistance (topical and intellectual discussions about the research which can lead to generation of new ideas)
- Affiliated institutions
- Grant numbers
- Who received the funding (if not the author, e.g. a supervisor)
- Any associated fellowships
Whom to acknowledge - and whom not to acknowledge
- Other authors/contributors : It is not common practice for the lead paper writer (i.e. the person writing and publishing the manuscript) to acknowledge the other authors/direct contributors to the paper. Only those who are not recognized as authors may be thanked and acknowledged.
- Reviewers : Authors are also not allowed to thank reviewers personally, or those who inspire them but cannot directly receive their appreciation – although reviewers can be thanked if they are kept anonymous .
- Friends and family : Unlike university manuscripts, journal manuscripts should not include help and guidance from family and friends.
Other acknowledgments
- Titles and institutions : Titles such as Mr, Mrs, Miss, etc. are not commonly included, but honorary titles such as Dr, Professor, etc. are. The institutions of the acknowledged people are usually mentioned.
For example, the following would not be acceptable:
We dedicate this work to the deceased Prof. Bloggs.
However, the following would be acceptable:
We acknowledge Prof. Bloggs for discovering the secret of anonymity.

Additional pointers for writing the Acknowledgments section
- The tone of the section should be in an active voice.
- Do not use pronouns indicating possession (i.e. his, her, their, etc.).
- Terms associated with specific companies should be written out in full, e.g. Limited, Corporation, etc.
- If the results have been published elsewhere, then this should also be acknowledged.
- Any abbreviations should be expanded unless the abbreviation appears in the main body of the text.
Below are examples of the Acknowledgments sections taken from a couple of papers from Nature Communications :

Duan L., Hope J., Ong Q., Lou H-Y., Kim N., McCarthy C., Acero V., Lin M., Cui B., Understanding CRY2 interactions for optical control of intracellular signalling, Nature Communications, 2017, 8:547
Xu Q., Jensen K., Boltyanskiy R., Safarti R., Style R., Dufresne E., Direct measurement of strain-dependent solid surface stress, Nature Communications, 2017, 8:555
Many people think that the Acknowledgments section of a manuscript is a trivial and unimportant component. However, it constitutes a vital means to ensure that all affiliated support for the paper can be duly and transparently mentioned. By acknowledging people for their efforts and contributions, you demonstrate your integrity as an academic researcher. In addition, crediting other people for their help can also increase their presence in the academic world and possibly help to boost their career as well as your own.
Maximise your publication success with Charlesworth Author Services.
Charlesworth Author Services, a trusted brand supporting the world’s leading academic publishers, institutions and authors since 1928.
To know more about our services, visit: Our Services
Share with your colleagues
Scientific Editing Services
Sign up – stay updated.
We use cookies to offer you a personalized experience. By continuing to use this website, you consent to the use of cookies in accordance with our Cookie Policy.

Acknowledgement for Thesis [Sample and Best Practice]
Advertisement
What is Acknowledgement in Thesis?
Writing a thesis is a significant milestone in a student’s academic journey. A vital component of a thesis is the acknowledgement section, which is often overlooked but holds immense importance. The acknowledgement section is a heartfelt expression of gratitude and recognition towards the individuals and organizations that have contributed to the completion of the thesis.
What is the purpose of acknowledgment and why it is important for thesis?
The purpose of the acknowledgement section in a thesis is to show appreciation to those who have supported and assisted the student in their research and writing process. This section provides an opportunity for the student to express their gratitude towards their supervisor, committee members, advisors, peers, friends, and family who have played a role in their academic journey. It also allows the student to acknowledge any financial support, resources, or facilities provided by institutions or organizations.
Acknowledgement in a thesis serves as a gesture of respect and gratitude towards those who have contributed to the student’s success. It is a way of recognizing the support and guidance received throughout the research and writing process. By acknowledging the contributions of others, the student acknowledges the collective effort that has gone into the completion of the thesis.
The importance of the acknowledgement section in a thesis cannot be understated. It is a way of paying tribute to the individuals and organizations that have supported and mentored the student, and it also reflects the student’s character and humility.
Additionally, it serves as a record of the student’s network and support system during their academic pursuit, and can also serve as a means of building and maintaining professional relationships in the academic community.
How to write best acknowledgement for thesis?
Here are some tips to write an effective acknowledgement for your thesis:
- Be Genuine and Sincere : Your acknowledgments should reflect genuine gratitude towards those who have contributed to your research journey. Avoid generic or insincere expressions of thanks.
- Personalize Your Thanks : Tailor your acknowledgments to acknowledge each individual’s specific contributions and support. Highlight unique experiences or insights they provided.
- Include Key Contributors : Acknowledge individuals who have directly contributed to your research, such as supervisors, mentors, collaborators, and participants. This could include academic advisors, industry professionals, field experts, or research assistants.
- Specify Contributions : Clearly articulate the contributions of each individual or group you’re acknowledging. Whether it’s providing guidance, sharing expertise, offering logistical support, or participating in discussions, specify how each person has helped you.
- Maintain Professionalism : While acknowledgments can be personal, maintain a professional tone and avoid overly informal language or inside jokes. Remember, this section is part of your academic work.
- Express Gratitude to Institutions : Acknowledge any institutions, organizations, or funding agencies that supported your research, providing grants, scholarships, or access to resources.
- Consider Cultural Sensitivities : Be mindful of cultural norms and sensitivities when expressing gratitude. Tailor your acknowledgments to reflect cultural expectations or customs, as appropriate.
- Proofread Carefully : Ensure your acknowledgments are free of grammatical errors and typos. Take the time to review and edit this section to ensure clarity and coherence.
- Maintain Balance : While it’s important to acknowledge those who have contributed significantly to your research, avoid overwhelming your acknowledgments with a long list of names. Prioritize key contributors and groups while being respectful of space limitations.
- Conclude with a General Expression of Thanks : After acknowledging specific individuals and groups, conclude your acknowledgments with a general expression of gratitude to anyone else who supported you in your academic journey. This could include family, friends, colleagues, or anyone else who provided encouragement or assistance.
Similar Guide: Writing acknowledgement best practice
Best Acknowledgement for Thesis
I extend my heartfelt gratitude to Dr. Maya Singh, Professor, and Chair of the Department of Ecology and Conservation Biology for granting me the opportunity to pursue this research endeavor.
I am indebted to my mentors, Professor Dr. Arjun Sharma from the Department of Ecology and Conservation Biology, for his unwavering guidance, invaluable advice, and relentless support throughout the course of this study.
Special thanks are due to Dr. Priya Patel, CEO of Environmental Solutions Ltd., for her generous provision of logistical support and invaluable insights, which greatly facilitated the successful execution of my research.
I express my deepest appreciation to Mr. Rajiv Gupta, Environmental Scientist at Environmental Solutions Ltd., Mr. Ankit Joshi, Environmental Engineer at Environmental Solutions Ltd., Ms. Priyanka Sharma, and Mr. Kevin Chen, Research Assistant at Stanford University, for their continuous assistance and support. Without their dedication and cooperation, this study would not have been possible.
I am also grateful to the entire team at Environmental Solutions Ltd. for their cooperation and assistance during the data collection process.
Special thanks go to Dr. Rahul Verma, Associate Professor at Delhi University, and Ms. Neha Kapoor, Environmental Consultant at Green Earth Solutions, for their encouragement and guidance throughout this project.
I would also like to extend my appreciation to my classmates from the Environmental Science program, as well as my friends, Rahul Singh, Ananya Mishra, and Deepak Sharma, for their unwavering support and encouragement throughout this journey.
Finally, I would like to express my deepest gratitude to my family, especially my elder brother Rohit, for his unwavering support and encouragement throughout my academic pursuits. His love and encouragement have been my greatest source of strength.

Acknowledgements for Thesis Example
I am profoundly grateful to the generous collaboration of numerous individuals whose collective efforts, wisdom, and warmth have imbued this thesis with depth and significance. Each person has left an indelible mark on this journey, contributing their unique perspectives and unwavering support. With deep gratitude, I extend my heartfelt acknowledgments, hopeful that this thesis may serve the cause of sustainable rural development in Madhya Pradesh.
First and foremost, I extend my sincere appreciation to the resilient farmers of Madhya Pradesh, whose resilience, patience, and hospitality have been the bedrock of this study. Their insights and experiences have been invaluable, shaping the very essence and direction of my research.
I owe a debt of gratitude to Mohan Patel for his pivotal suggestion to connect with Dr. Anjali Deshmukh and pursue an internship at the Madhya Pradesh Rural Development Institute (MPRDI). I am immensely thankful to Dr. Anjali Deshmukh, my mentor, whose guidance and mentorship have illuminated my path in rural development, offering invaluable advice and enlightening discussions.
Special thanks are extended to Rajesh Singh for his friendship, hospitality, and profound conversations, as well as for providing me with the opportunity to present my findings at MPRDI. I also extend my gratitude to Prakash Gupta for his expertise, passion, and invaluable guidance, and to Rani Devi for her companionship in the field and insightful discussions.
I express my heartfelt appreciation to Om Prakash, Mukesh Kumar, Sunita Devi, Ramesh Patel, and Kamla Bai for generously sharing their knowledge, engaging in insightful discussions, and offering invaluable advice. My gratitude also goes to Raju Verma, Rameshwar Singh, Vijay Kumar, Deepak Yadav, and Babita Devi for their kindness, sharing their invaluable field experiences, and the cherished moments we spent together.
Special thanks are due to Rajesh Sharma, Rajni Devi, Nirmala Bai, and Ravi Kumar for their assistance, which significantly enriched my understanding of local practices and challenges. I also extend my gratitude to Shanti Devi and Santosh Kumar for their unwavering support and guidance throughout my fieldwork in MPRDI.
Heartfelt thanks are extended to Vikas, Sunil, and Anil for their companionship during our travels across rural communities, driven by their passion for positive change. I also express my gratitude to Kamlesh Devi for our enriching discussions, camaraderie, and friendship.
Furthermore, I extend my appreciation to all participants of the seminars in Bhopal, whose generosity in allowing me to share my findings and providing invaluable feedback has been invaluable. I am particularly grateful to everyone who facilitated the dialogue between MPRDI and local stakeholders regarding the results presented in this thesis.
Note: All names and institutions mentioned here are fictional and used for sample purposes only.
Related:
- Acknowledgement for Dissertation
- Acknowledgement in Research Paper
Best Practice
In addition to listing contributions, briefly mention how each person’s support or expertise impacted your research or personal growth. This adds depth to your acknowledgments and shows genuine appreciation for their role in your academic journey. For example, you could mention specific insights gained from discussions, how their guidance shaped your methodology, or how their support bolstered your confidence during challenging times. This not only highlights the significance of their contributions but also reinforces the depth of your gratitude.
Similar Posts
Acknowledgement for commerce project pdf.
Advertisement Commerce project for Class 12 students is a significant academic endeavor that requires extensive analysis, and compilation…
Acknowledgement for Portfolio | How to Write | With Sample
Advertisement Acknowledgement for portfolio is crucial for recognizing the contributions and efforts of others. It serves as a…
Acknowledgement for Dissertation [With Example]
Advertisement Writing a dissertation is a significant academic undertaking that requires extensive research, critical analysis, and original contributions…
Acknowledgement for Group Project [PDF Sample]
Advertisement When it comes to group projects, acknowledging the individual contributions of each team member is not just…
Acknowledgement for Assignment (6 Sampels with PDF)
Advertisement Acknowledgement is a pivotal aspect of any assignment, showcasing gratitude towards those who have contributed to its…
Acknowledgement in Research Paper | How to Write | Perfect Example
Advertisement What is acknowledgement in research paper? Acknowledgement in a research paper is the section where the author…
Academia Bees
Acknowledgement for Paper Publication (10 Samples)
July 11, 2023
No Comments
By Mohsin Khurshid
Acknowledgement for paper publication is a crucial aspect that recognizes the contributions and support received throughout the research process. It holds significant value in academic papers, showcasing appreciation for the individuals and institutions that have played a role in the study’s success. In this article, we will delve into the importance of acknowledgement in paper publication and provide 10 comprehensive samples to guide you in crafting impactful acknowledgements for your own research. Whether you wish to acknowledge research collaborators, funding sources, mentors, or other individuals who have contributed to your work, these samples will assist you in expressing your gratitude effectively. Join us as we explore the art of acknowledgement and discover how to create acknowledgements that resonate with the collaborative nature of scholarly endeavors.
Table of Contents
- 1 Understanding the Importance of Acknowledgement
- 2.1 Sample 1 – Acknowledgement for Research Collaboration
- 2.2 Sample 2 – Acknowledgement for Funding Support
- 2.3 Sample 3 – Acknowledgement for Mentorship and Guidance
- 2.4 Sample 4 – Acknowledgement for Institutional Support
- 2.5 Sample 5 – Acknowledgement for Peer Reviewers
- 2.6 Sample 6 – Acknowledgement for Data and Materials
- 2.7 Sample 7 – Acknowledgement for Technical Assistance
- 2.8 Sample 8 – Acknowledgement for Editorial or Proofreading Assistance
- 2.9 Sample 9 – Acknowledgement for Inspiration and Support
- 2.10 Sample 10 – Acknowledgement for Personal Support
- 4 Conclusion
Understanding the Importance of Acknowledgement
Acknowledgement plays a crucial role in academic papers, serving multiple purposes within the scholarly community. Firstly, it provides an opportunity for authors to express gratitude and appreciation towards individuals, organizations, or institutions that have contributed to the research project in various ways. These contributions can include financial support, data collection assistance, intellectual guidance, or access to resources.
Beyond mere gratitude, acknowledgement serves as a means of giving credit where it is due. It recognizes the valuable input and collaboration of others, ensuring that their contributions are acknowledged and acknowledged by the academic community. By acknowledging the efforts of others, researchers uphold the principles of academic integrity and promote transparency in the research process.
Ethical considerations are also an integral part of writing an acknowledgement section. Authors must carefully navigate issues such as conflicts of interest, ensuring that they disclose any potential conflicts and maintain objectivity in their acknowledgements. Additionally, it is essential to respect the privacy and confidentiality of individuals mentioned in the acknowledgement, seeking their permission before including their names or affiliations.
Conventions related to acknowledgement may vary across disciplines and academic journals. Authors should familiarize themselves with the specific guidelines and requirements of their target journal to ensure compliance. Some journals may provide templates or specific instructions on how to structure the acknowledgement section, while others may have specific word limits or content restrictions.
In summary, understanding the importance of acknowledgement in academic papers is crucial for researchers. It serves as a means of expressing gratitude, giving credit to contributors, upholding academic integrity, and adhering to ethical considerations. By following conventions and guidelines specific to their field, authors can effectively convey their appreciation and recognition in the acknowledgement section of their paper.
10 Samples of Acknowledgement for Paper Publication
Explore these detailed samples of acknowledgements to effectively acknowledge contributors in your research papers.
Sample 1 – Acknowledgement for Research Collaboration
Acknowledging the collaborative efforts and contributions of research team members is a crucial aspect of the acknowledgement section in academic papers. The following is a detailed sample acknowledgement that demonstrates how to acknowledge research collaborators and team members:
I would like to express my sincere gratitude to the members of the research team who contributed to the successful completion of this study. Their dedication, expertise, and commitment were instrumental in the realization of our research objectives. I am thankful for their valuable insights, collaborative spirit, and unwavering support throughout the project.
I would like to extend my heartfelt appreciation to [Name], [Name], and [Name] for their invaluable contributions to the data collection process. Their meticulous efforts ensured the accuracy and reliability of our research findings. Additionally, I am grateful to [Name] for their assistance in data analysis and interpretation, which greatly enhanced the depth and quality of our research outcomes.
Furthermore, I would like to acknowledge the guidance and mentorship provided by [Name]. Their expertise in [area of expertise] was invaluable in shaping the direction of this research and refining our methodology. Their constructive feedback and insightful suggestions played a pivotal role in elevating the overall quality of this study.
I am also indebted to the support staff and administrators who facilitated the smooth operation of our research activities. Their assistance in securing necessary permissions, organizing logistics, and managing administrative tasks was vital to the success of this project.
Lastly, I would like to express my deep appreciation to the funding agency or organization that provided financial support for this research. Their investment in our work enabled us to conduct this study and make meaningful contributions to the field.
In conclusion, I am immensely grateful to all the individuals who contributed to this research collaboration. Their dedication, expertise, and unwavering support have been instrumental in the accomplishment of our research goals. Without their invaluable contributions, this study would not have been possible.
Sample 2 – Acknowledgement for Funding Support
Acknowledging the financial support received for research is an important aspect of the acknowledgement section in academic papers. The following is a detailed sample acknowledgement that demonstrates how to acknowledge funding support:
I would like to express my sincere appreciation to [Funding Agency/Organization] for their generous financial support of this research project. Their funding played a crucial role in the successful execution of this study and the attainment of our research goals.
The support provided by [Funding Agency/Organization] enabled us to conduct data collection, analysis, and interpretation, as well as cover expenses related to research materials, participant recruitment, and travel, where applicable. Their investment in our work has significantly contributed to the quality and impact of our research findings.
I would also like to extend my gratitude to the grant administrators and program officers at [Funding Agency/Organization] for their guidance and assistance throughout the grant application and management process. Their expertise and support were invaluable in ensuring a smooth and efficient funding experience.
Furthermore, I am grateful to my research team members and collaborators who have contributed their time, expertise, and efforts to this project. Their dedication and hard work have been instrumental in the successful completion of this research.
In conclusion, I am deeply thankful to [Funding Agency/Organization] for their financial support, without which this research would not have been possible. Their investment in our work has made a significant impact and has contributed to advancements in the field. I am truly grateful for their commitment to supporting research and fostering academic growth.
Sample 3 – Acknowledgement for Mentorship and Guidance
Acknowledging the contributions of mentors and advisors is essential in recognizing the guidance and support they have provided throughout the research process. The following is a detailed sample acknowledgement that demonstrates how to acknowledge mentorship and guidance:
I would like to express my heartfelt gratitude to my mentor, [Mentor’s Name], for their invaluable guidance, expertise, and support throughout the course of this research. Their extensive knowledge, insightful feedback, and unwavering encouragement have been instrumental in shaping the direction and quality of this study.
I am truly grateful for [Mentor’s Name]’s dedication and commitment to my academic growth. Their mentorship has not only expanded my understanding of the subject matter but has also enhanced my research skills and critical thinking abilities. Their willingness to invest their time and effort in providing guidance and constructive criticism has greatly contributed to the successful completion of this research project.
I would also like to extend my appreciation to [Advisor’s Name] for their valuable input and advice throughout the research process. Their expertise in [specific field or area] has been invaluable in shaping the methodology and interpretation of the findings. Their constructive feedback and thoughtful suggestions have significantly enhanced the quality and rigor of this study.
Furthermore, I would like to acknowledge the contributions of other members of my research committee, [Committee Member 1’s Name] and [Committee Member 2’s Name]. Their expertise and insights have provided valuable perspectives and have contributed to the overall excellence of this research.
In conclusion, I am sincerely grateful to my mentor, [Mentor’s Name], and my advisor, [Advisor’s Name], for their unwavering support and guidance. Their mentorship and expertise have been transformative in my academic journey, and I am privileged to have had the opportunity to learn from their wisdom and experience.
Sample 4 – Acknowledgement for Institutional Support
Acknowledging the support and resources provided by institutions is crucial in recognizing their contribution to the research project. The following is a detailed sample acknowledgement that demonstrates how to acknowledge institutional support:
I would like to express my sincere gratitude to [Institution’s Name] for their unwavering support and provision of resources throughout the course of this research. The research facilities and infrastructure provided by [Institution’s Name] have played a significant role in the successful completion of this study.
I am grateful for the access to state-of-the-art laboratories, research materials, and technological resources offered by [Institution’s Name]. These resources have been instrumental in conducting experiments, gathering data, and analyzing findings. The research environment provided by [Institution’s Name] has fostered an atmosphere of innovation and collaboration, allowing for the pursuit of academic excellence.
I would also like to extend my appreciation to the administrative staff and personnel at [Institution’s Name] for their assistance and support throughout the research process. Their responsiveness and willingness to provide guidance on administrative matters have been invaluable.
Furthermore, I would like to acknowledge the financial support provided by [Institution’s Name]. The research grants and scholarships awarded by [Institution’s Name] have enabled the realization of this project and have alleviated the financial constraints associated with conducting research.
In conclusion, I am deeply grateful to [Institution’s Name] for their unwavering support, provision of resources, and financial assistance. Their commitment to promoting research and academic excellence has been instrumental in the successful completion of this study.
Sample 5 – Acknowledgement for Peer Reviewers
Acknowledging the contributions of peer reviewers is essential in recognizing their valuable feedback and input towards improving the quality of the research. The following is a detailed sample acknowledgement that demonstrates how to acknowledge peer reviewers:
I would like to extend my heartfelt appreciation to the anonymous peer reviewers who generously dedicated their time and expertise to review and provide constructive feedback on this research paper. Their insightful comments and suggestions have significantly enhanced the quality and rigor of this study.
The meticulous review process conducted by the peer reviewers has played a crucial role in strengthening the methodology, refining the arguments, and improving the overall clarity and coherence of the research. Their expertise and critical evaluation have helped in identifying and addressing potential gaps, ensuring the accuracy and validity of the findings.
I am grateful for the valuable contributions made by the peer reviewers in shaping this paper into its final form. Their commitment to scholarly excellence and their dedication to advancing the field have been instrumental in improving the quality and impact of this research.
While their identities remain anonymous, I would like to express my sincere gratitude to each and every peer reviewer who played a part in the review process. Their time, expertise, and feedback are deeply appreciated.
Sample 6 – Acknowledgement for Data and Materials
Acknowledging the sources of data, materials, or tools used in the research is crucial to give proper credit and recognition. The following is a detailed sample acknowledgement that demonstrates how to acknowledge data and materials:
I would like to express my sincere gratitude to [Name/Organization] for providing access to the [Specify the data/materials/tools] used in this research. Their contribution has been invaluable in facilitating the data collection process and enabling the analysis conducted in this study.
The [Specify the data/materials/tools] have played a pivotal role in the success of this research, providing essential insights, information, and resources that have contributed to the depth and quality of the findings. Without their support and cooperation, this research would not have been possible.
I am deeply appreciative of the efforts made by [Name/Organization] in making the [Specify the data/materials/tools] available and ensuring their reliability and relevance to the research objectives. Their commitment to data sharing and collaborative research has greatly enriched the outcomes of this study.
I would also like to extend my gratitude to the individuals involved in the collection, organization, and maintenance of the [Specify the data/materials/tools]. Their dedication and professionalism have been instrumental in making these resources accessible and usable for this research.
Sample 7 – Acknowledgement for Technical Assistance
Acknowledging the individuals or organizations that provided technical assistance is essential to recognize their contributions. The following is a detailed sample acknowledgement that demonstrates how to acknowledge technical support:
I would like to extend my sincere appreciation to [Name/Organization] for their invaluable technical assistance throughout the course of this research. Their expertise and support have played a crucial role in the successful execution of this project.
The technical assistance provided by [Name/Organization] has been instrumental in overcoming various challenges and ensuring the accuracy and reliability of the experimental procedures. Their guidance and knowledge have greatly enhanced the quality of the data collected and the interpretation of the results.
I am grateful for their willingness to share their expertise and resources, providing guidance on experimental techniques, troubleshooting technical issues, and offering valuable insights into data analysis. Their assistance has significantly contributed to the overall success of this research endeavor.
I would also like to express my gratitude to the individuals within [Name/Organization] who have directly contributed to this project. Their dedication, professionalism, and prompt response to queries have made the research process smoother and more efficient.
Sample 8 – Acknowledgement for Editorial or Proofreading Assistance
Acknowledging the individuals who have contributed to the editing and proofreading process is essential to recognize their valuable contributions. The following is a detailed sample acknowledgement that demonstrates how to acknowledge editorial or proofreading assistance:
I would like to express my sincere gratitude to [Name/Names] for their invaluable assistance in the editing and proofreading of this manuscript. Their keen attention to detail, expertise in language and grammar, and commitment to improving the clarity and readability of the content have significantly enhanced the quality of this work.
The diligent efforts of [Name/Names] in meticulously reviewing the manuscript, suggesting revisions, and providing valuable feedback have played a crucial role in refining the overall structure, organization, and coherence of the document. Their editorial skills have helped to ensure the accuracy and precision of the scientific content, enhancing the credibility and impact of the research findings.
I am deeply appreciative of their dedication and professionalism throughout the editing process. Their constructive criticism, insightful suggestions, and meticulous proofreading have greatly contributed to the refinement of the language, grammar, and style of this manuscript.
I would also like to extend my thanks to [Name/Names] for their timely and efficient collaboration, as well as their willingness to accommodate multiple rounds of revisions. Their commitment to excellence and their commitment to producing a polished final product are greatly appreciated.
Sample 9 – Acknowledgement for Inspiration and Support
Acknowledging the individuals or sources of inspiration that have contributed to the research is important in recognizing their impact on the project. The following is a detailed sample acknowledgement that demonstrates how to acknowledge inspiration and support:
I would like to express my heartfelt gratitude to [Name/Names] for their unwavering support and inspiration throughout this research endeavor. Their encouragement, guidance, and insightful discussions have been instrumental in shaping the direction and depth of this study.
The unwavering support of [Name/Names] has been a constant source of motivation, providing the necessary encouragement to overcome challenges and pursue innovative ideas. Their expertise, wisdom, and constructive feedback have played a pivotal role in refining the research objectives, methodology, and interpretation of findings.
I am also indebted to [Source/Event/Book/Quote] for serving as a source of inspiration and influencing the conceptual framework of this study. The [Source/Event/Book/Quote] has provided valuable insights and perspectives that have guided my thinking and contributed to the overall development of this research.
Additionally, I would like to acknowledge the support and understanding of my family, friends, and colleagues, whose unwavering belief in my abilities has sustained me throughout this research journey. Their encouragement, understanding, and patience have been invaluable, providing the necessary emotional support and creating an environment conducive to pursuing this endeavor.
Sample 10 – Acknowledgement for Personal Support
Acknowledging the emotional support provided by family, friends, and loved ones is essential in recognizing their role in the research process. The following is a detailed sample acknowledgement that demonstrates how to acknowledge personal support:
I would like to express my deepest appreciation to my family, friends, and loved ones for their unwavering support throughout this research journey. Their love, understanding, and encouragement have been the pillars that have kept me motivated and focused during challenging times.
To my [Family Member(s)], your unwavering belief in me and your constant encouragement have been the driving force behind my pursuit of this research. Your sacrifices, understanding, and patience have provided me with the necessary space and time to dedicate myself to this endeavor. I am forever grateful for your unconditional love and support.
I would also like to extend my gratitude to my dear friends who have been a source of inspiration and motivation. Your unwavering belief in my abilities, your listening ear, and your words of encouragement have given me the strength and confidence to overcome obstacles and pursue this research with dedication.
In addition, I am grateful to [Name(s)] for their continuous support, understanding, and patience. Their presence and unwavering support have provided me with the emotional stability and reassurance needed to navigate through the challenges of this research journey.
Finally, I want to express my heartfelt appreciation to all the participants who generously shared their time, experiences, and insights for this research. Without their willingness to contribute, this study would not have been possible.

Also, read tips and samples on writing acknowledgement for your internship report .
Writing an acknowledgement for a paper involves expressing gratitude to individuals or entities who have contributed to the research. It typically includes mentioning the names, roles, and specific contributions of those being acknowledged.
To acknowledge means to recognize and show appreciation for the contributions or support received. In the context of paper publication, it refers to acknowledging the individuals or organizations that have played a role in the research process.
In an acknowledgement section, you can express gratitude to individuals, institutions, or funding agencies that have supported or contributed to the research. You can mention their names, roles, and specific contributions in a sincere and respectful manner.
The correct spelling of “acknowledge” is A-C-K-N-O-W-L-E-D-G-E.
An acknowledgement statement refers to a written expression of gratitude and appreciation towards individuals or organizations for their contributions or support. It is a way to formally recognize their involvement in the research process and show gratitude for their assistance.
In conclusion, acknowledgement plays a vital role in paper publication, allowing researchers to express gratitude and recognize the contributions of individuals and institutions who have supported their work. By acknowledging research collaborators, mentors, funding sources, peer reviewers, and others, researchers demonstrate their appreciation and foster a sense of community within the academic sphere.
It is important to tailor acknowledgements to the specific research context, considering the unique contributions and support received throughout the research process. Each acknowledgement should reflect the genuine appreciation and recognition of those who have played a significant role in the success of the research.
As researchers, let us not underestimate the power of acknowledgement. It not only acknowledges the contributions of others but also serves as a reminder of the collaborative and interconnected nature of academic work. By expressing gratitude and recognizing the efforts of those who have supported us, we contribute to a positive research culture and encourage further collaboration and support in the academic community . In closing, let us remember that acknowledgement is not just a formality but a sincere gesture of appreciation. It is a way to honor the collaborative nature of research and the individuals and institutions that have contributed to our academic journey. May we continue to acknowledge and support one another as we advance knowledge and make meaningful contributions to our respective fields.
Rules for Dealing with Introverts: Complete Guide
How to write acknowledgement for research paper (5 samples), leave a comment cancel reply.
Save my name, email, and website in this browser for the next time I comment.
Acknowledgement Letter
Get All Types Of Acknowledgement Samples.
How To Write Acknowledgement For Research Paper
Post by Ruben Patel Leave a Comment
If you are thinking about How To Write Acknowledgment For Research Paper then you have come to the right place.
An acknowledgement section in a research paper is a place where you can express your gratitude to the people who have helped you with your research. This could include anyone who has provided support , guidance , or assistance during the research process.
For example, you might thank your advisors or professors for their guidance, your colleagues for their collaboration, librarians, lab assistants or a funding agency for their financial support and anyone else who has contributed to your research.
It is common to include an acknowledgement section in the beginning of a research paper, usually after the abstract and before the main body of the paper. The acknowledgement should be brief and to the point, and it is generally written in a formal, professional tone. You should avoid using casual language or personal anecdotes in this section.

The purpose of an acknowledgement is to express your gratitude to the people who have helped you with your research. It is a way to show that you appreciate the support and contributions of others, and it is an opportunity to thank them for their assistance. By including an acknowledgement in your research paper, you demonstrate that you are grateful for the help and support you have received, and you recognize the role that others have played in your research.
Here are some tips for writing an acknowledgement for a research paper:
- Start by thanking the people who have directly supported your research, such as your advisors and colleagues.
- Mention any funding agencies or organizations that provided financial support for your research.
- If you received assistance from librarians or other research staff, be sure to thank them as well.
- If you received help from any other individuals, such as lab assistants or research participants, be sure to include them in your acknowledgement.
- Keep your acknowledgement brief and to the point. You don’t need to go into great detail about the contributions of each person you are thanking.
- Use a formal, professional tone in your acknowledgement. This is not the place for casual language or personal anecdotes.
Here are a few acknowledgement samples for your research paper –
Table of Contents
Acknowledgement For Research Paper Sample
We are deeply grateful to all those who contributed to the success of this research project.
First and foremost, we would like to thank our primary supervisor, [Name], for their guidance, support, and encouragement throughout the entire process. Their mentorship and expertise were invaluable in helping us to shape the direction of our research and to bring our ideas to fruition.
We would also like to express our gratitude to the members of our research team, [Names], who provided valuable input, insights, and assistance at every stage of the project. Their contributions were critical to the success of this research, and we are deeply grateful for their hard work and dedication.
We would also like to thank the organizations and individuals who provided financial support for this research, including [Name] and [Name]. Without their generous contributions, this project would not have been possible.
Finally, we would like to extend our heartfelt thanks to all of the participants in our study, who generously shared their time, experiences, and insights with us. Their willingness to engage with our research was essential to the success of this project, and we are deeply grateful for their participation.
Overall, this research project would not have been possible without the support and contributions of so many people. We are deeply grateful to all of those who helped to make this project a reality, and we hope that our findings will make a meaningful contribution to the field.
Read Also: Get 15+ Acknowledgement Samples
Sample Acknowledgement For Research Paper
The completion of this research project would not have been possible without the contributions and support of many individuals and organizations. We are deeply grateful to all those who played a role in the success of this project.
We would like to thank [Name] for their invaluable input and support throughout the research process. Their insights and expertise were instrumental in shaping the direction of this project.
In addition, we would like to extend our sincere thanks to all of the participants in our study, who generously shared their time, experiences, and insights with us. Their willingness to engage with our research was essential to the success of this project, and we are deeply grateful for their participation.
Small Acknowledgement Sample For Research Paper
We would like to express our sincere gratitude to our advisors, Dr. Spirt Kular and Dr. Sonney Deom, for their invaluable guidance and support throughout the research process. We also wish to thank the XYZ Foundation for their financial support, as well as the librarians at the ABC Library for their assistance in finding the necessary research materials. Finally, we are grateful to all of the research participants who generously gave their time and effort to this project.
You can take the help online paraphrasing tool when crafting an acknowledgment for a research paper. They help you manipulate the formal tone of your text according to your needs and make sure your text stays sincere and relevant.
Using the paraphrase tool , you can express specific appreciation for individual contributions that fit the unique context of the paper. Ultimately, you write an acknowledgment that is as personalized and meaningful as possible.
Note: It’s important to note that the specific language and tone of your acknowledgement will depend on the context and the nature of the contributions made by the individuals or organizations you are thanking. It’s a good idea to be specific and sincere in your appreciation, and to tailor your acknowledgement to the specific contributions that were made.
Conclusion On How To Write Acknowledgement For Research Paper
In conclusion, writing an acknowledgement for a research paper is a crucial step in recognizing the contributions of others to your work. It is a way to show gratitude to those who have provided assistance, support, and guidance throughout the research process. When writing an acknowledgement, it is important to be sincere and specific in thanking those who have helped you. Be sure to mention individuals by name, and specify their roles and contributions.
Additionally, make sure to follow any specific guidelines or requirements for formatting and presenting the acknowledgement in your research paper. By following these tips, you can effectively convey your appreciation and recognition of the help you received while completing your research.
Read Also: Master Thesis Acknowledgement Sample
FAQs On How To Write Acknowledgement For Research Paper
1) What is a good sentence for acknowledgement? Ans: “I appreciate your efforts in completing this project on time.” “I want to acknowledge the hard work and dedication you have shown during this challenging year.” “I am grateful for your support and understanding during this difficult time.” It’s important to note that the specific wording and tone of an acknowledgement will depend on the context and the relationship between the speaker and the person being acknowledged.
2) Is conclusion and acknowledgement same? Ans: The conclusion is the final part of a written work that summarizes the main points and arguments. Acknowledgement is the act of recognizing or thanking someone or something for their contribution. These two things are often used in different contexts and for different purposes.
3) Who should I include in my acknowledgement for a research paper? Ans: It is appropriate to include anyone who has contributed significantly to your research, such as a supervisor, mentor, or colleague. You may also want to thank any individuals or organizations that provided financial or logistical support.
4) How should I structure my acknowledgement for a research paper? Ans: The structure of your acknowledgement will depend on the specific guidelines of the journal or publication you are submitting to. In general, it is common to include a brief paragraph at the beginning of the paper expressing your gratitude to those who have helped with the research. You can then list the names of the individuals or organizations you are thanking, along with a brief description of their contributions.
5) Is it necessary to include an acknowledgement in my research paper? Ans: It is not always required to include an acknowledgement in a research paper, but it is a common practice and can be a good way to show appreciation for the help and support you received during the research process.
6) Can I include personal thanks in my acknowledgement for a research paper? Ans: It is generally acceptable to include personal thanks in your acknowledgement, as long as it is done in a professional and respectful manner. You may want to thank family members or friends who provided emotional support during the research process, but be mindful of maintaining a professional tone and not going into too much detail about personal matters.
Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.

- Skip to main content
- Skip to search
- Skip to footer
Products and Services

Cisco Provider Connectivity Assurance
Gain end-to-end visibility and insights like never before..
Create exceptional digital experiences built on deep network observability and critical network monitoring.
Service assurance that's proactive and precise—a win-win
Drive additional sales revenue.
See more revenue from portal fees and capacity purchases when you give customers deeper visibility and transparency with Cisco Provider Connectivity Assurance.
Boost efficiency and revenue with Cisco Provider Connectivity Assurance (formerly Accedian Skylight), delivering service assurance that continuously monitors and optimizes digital experiences.
Simplified operations and troubleshooting
Gain a single view of granular performance metrics and third-party data to accelerate MTTR.
Continuous optimization of digital experiences
Find transient issues with precise synthetic network and service testing.
Differentiated services, next-level innovation
Premium, SLA-backed services and end-customer portals are just a click away.
Predictive analytics powered by AI
Be proactive, not reactive. Put the power of your multivendor infrastructure to work for a trouble-free network.
See what sets Cisco Provider Connectivity Assurance apart
Granular, real-time visibility.
Perform end-to-end and full mesh testing with microsecond precision and make hidden microbursts a thing of the past.
Reliability meets flexibility
Enjoy 99.997% uptime and take full control over network performance for greater reliability.
Optimize your infrastructure
Automatically correlate sensor metrics with third-party sources of performance data to improve ROI and create better digital experiences.
Deliver exceptional services
Ready-made customer portals differentiate services, boost revenue, and enhance satisfaction.
Assuring high-performance critical networks
See how Cisco Provider Connectivity Assurance enables efficient troubleshooting and SLA assurance—all while lowering operational costs.
Cisco Provider Connectivity Assurance use cases
Solve the challenges of fragmented multidomain tools and poor visibility on service quality. Discover how Provider Connectivity Assurance can help you simplify operations.
Real-time SLA visibility
Monitor and police SLAs for accountability, while proactively addressing performance issues.
Critical network monitoring
Improve operational efficiency with proactive assurance and microsecond visibility for faster issue resolution and better experiences.
B2B service differentiation
Create new revenue opportunities to upsell and differentiate your services with multitenant end-customer portals for real-time service-level agreement (SLA) visibility and alerting.
Automated assurance
Detect issues early when you automate assurance for the entire service lifecycle with real-time service visibility and predictive analytics.
Multilayer assurance
Get a single view of performance across multiple network layers, like segment routing and routed optical networking, to reduce tools and drive down MTTR.
Mobile backhaul and edge monitoring
Optimize digital experiences with real-time service visibility, while assuring your end-to-end 5G transport.
Get high-precision service assurance with Cisco Provider Connectivity Assurance
Ai-enabled predictive analytics.
Get an ML-assisted view of your entire network and gain powerful performance insights.
Assurance sensors
Network-wide service performance visibility. Deployable anywhere, at scale.
See why your peers trust Cisco Provider Connectivity Assurance
"we can look at network performance at any level.".
With Cisco Provider Connectivity Assurance, Bouygues Telecom now has a complete "telescopic and microscopic" view of network and service performance in a single tool.
André Ethier, Network Quality Engineer
Bouygues Telecom
"The solution is key in evolving our end-to-end network visibility."
This is extremely powerful in terms of customer experience. It helps to avoid tickets, as customers can see for themselves what happened with their service. This reduces tension and increases customer satisfaction.
Bart Janssens, Senior Specialist, Packet Architecture
Take preventative measures against network degradation.
"With service-centric assurance and granular visibility we can prevent degradations, automate actions for improvements, and better communicate with our customers."
Mahesh Anjan, Senior Product Technology Executive
Better together
Cisco crosswork network automation.
Drive network efficiency and enhance experiences.
Cisco Optics
Plug innovation into your network.
Boost efficiency and service quality
Cisco Provider Connectivity Assurance helps you improve service quality, lower costs, and deliver outstanding user experiences with a single view of service performance across the entire network
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, automatically generate references for free.
- Knowledge Base
- Dissertation
- Thesis & Dissertation Acknowledgements | Tips & Examples
Thesis & Dissertation Acknowledgements | Tips & Examples
Published on 4 May 2022 by Tegan George . Revised on 4 November 2022.
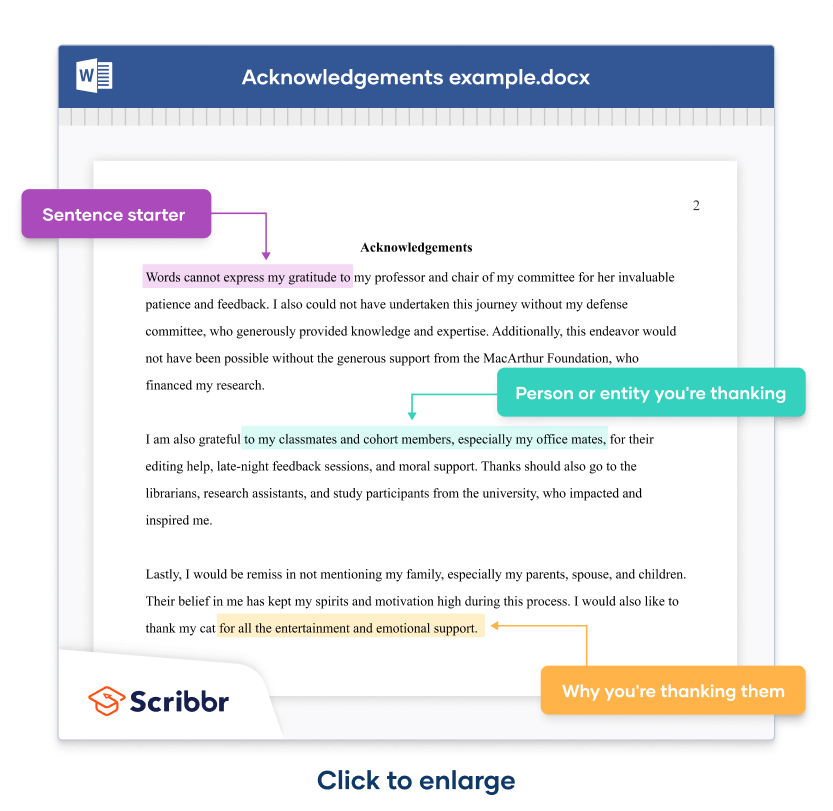
The acknowledgements section is your opportunity to thank those who have helped and supported you personally and professionally during your thesis or dissertation process.
Thesis or dissertation acknowledgements appear between your title page and abstract and should be no longer than one page.
In your acknowledgements, it’s okay to use a more informal style than is usually permitted in academic writing , as well as first-person pronouns . Acknowledgements are not considered part of the academic work itself, but rather your chance to write something more personal.
To get started, download our step-by-step template in the format of your choice below. We’ve also included sample sentence starters to help you construct your acknowledgments section from scratch.
Download Word doc Download Google doc
Instantly correct all language mistakes in your text
Be assured that you'll submit flawless writing. Upload your document to correct all your mistakes.

Table of contents
Who to thank in your acknowledgements, how to write acknowledgements, acknowledgements section example, acknowledgements dos and don’ts, frequently asked questions.
Generally, there are two main categories of acknowledgements: professional and personal .
A good first step is to check your university’s guidelines, as they may have rules or preferences about the order, phrasing, or layout of acknowledgements. Some institutions prefer that you keep your acknowledgements strictly professional.
Regardless, it’s usually a good idea to place professional acknowledgements first, followed by any personal ones. You can then proceed by ranking who you’d like to thank from most formal to least.
- Chairs, supervisors, or defence committees
- Funding bodies
- Other academics (e.g., colleagues or cohort members)
- Editors or proofreaders
- Librarians, research/laboratory assistants, or study participants
- Family, friends, or pets
Typically, it’s only necessary to mention people who directly supported you during your thesis or dissertation. However, if you feel that someone like a secondary school physics teacher was a great inspiration on the path to your current research, feel free to include them as well.
Professional acknowledgements
It is crucial to avoid overlooking anyone who helped you professionally as you completed your thesis or dissertation. As a rule of thumb, anyone who directly contributed to your research should be mentioned.
A few things to keep in mind include:
- Even if you feel your chair didn’t help you very much, you should still thank them first to avoid looking like you’re snubbing them.
- Be sure to follow academic conventions, using full names with titles where appropriate.
- If several members of a group or organisation assisted you, mention the collective name only.
- Remember the ethical considerations around anonymised data. If you wish to protect someone’s privacy, use only their first name or a generic identifier (such as ‘the interviewees’).
Personal acknowledgements
There is no need to mention every member of your family or friend group. However, if someone was particularly inspiring or supportive, you may wish to mention them specifically. Many people choose to thank parents, partners, children, friends, and even pets, but you can mention anyone who offered moral support or encouragement, or helped you in a tangible or intangible way.
Some students may wish to dedicate their dissertation to a deceased influential person in their personal life. In this case, it’s okay to mention them first, before any professional acknowledgements.
The only proofreading tool specialized in correcting academic writing
The academic proofreading tool has been trained on 1000s of academic texts and by native English editors. Making it the most accurate and reliable proofreading tool for students.

Correct my document today
After you’ve compiled a list of who you’d like to thank, you can then sort your list into rank order. Separate everyone you listed into ‘major thanks’, ‘big thanks’, and ‘minor thanks’ categories.
- ‘Major thanks’ are given to people who your project would be impossible without. These are often predominantly professional acknowledgements, such as your advisor , chair, and committee, as well as any funders.
- ‘Big thanks’ are an in-between, for those who helped you along the way or helped you grow intellectually, such as classmates, peers, or librarians.
- ‘Minor thanks’ can be a catch-all for everyone else, especially those who offered moral support or encouragement. This can include personal acknowledgements, such as parents, partners, children, friends, or even pets.
How to phrase your acknowledgements
To avoid acknowledgements that sound repetitive or dull, consider changing up your phrasing. Here are some examples of common sentence starters you can use for each category.
| Major thanks | Big thanks | Minor thanks |
|---|---|---|
Note that you do not need to write any sort of conclusion or summary at the end. You can simply end the acknowledgements with your last thank-you.
Here’s an example of how you can combine the different sentences to write your acknowledgements.
A simple construction consists of a sentence starter (in purple highlight ), followed by the person or entity mentioned (in green highlight ), followed by what you’re thanking them for (in yellow highlight .)
Acknowledgements
Words cannot express my gratitude to my professor and chair of my committee for her invaluable patience and feedback. I also could not have undertaken this journey without my defense committee, who generously provided knowledge and expertise. Additionally, this endeavor would not have been possible without the generous support from the MacArthur Foundation, who financed my research .
I am also grateful to my classmates and cohort members, especially my office mates, for their editing help, late-night feedback sessions, and moral support. Thanks should also go to the librarians, research assistants, and study participants from the university, who impacted and inspired me.
Lastly, I would be remiss in not mentioning my family, especially my parents, spouse, and children. Their belief in me has kept my spirits and motivation high during this process. I would also like to thank my cat for all the entertainment and emotional support.
- Write in first-person, professional language
- Thank your professional contacts first
- Include full names, titles, and roles of professional acknowledgements
- Include personal or intangible supporters, like friends, family, or even pets
- Mention funding bodies and what they funded
- Appropriately anonymise or group research participants or non-individual acknowledgments
Don’t:
- Use informal language or slang
- Go over one page in length
- Mention people who had only a peripheral or minor impact on your work
Prevent plagiarism, run a free check.
You may acknowledge God in your thesis or dissertation acknowledgements , but be sure to follow academic convention by also thanking the relevant members of academia, as well as family, colleagues, and friends who helped you.
Yes, it’s important to thank your supervisor(s) in the acknowledgements section of your thesis or dissertation .
Even if you feel your supervisor did not contribute greatly to the final product, you still should acknowledge them, if only for a very brief thank you. If you do not include your supervisor, it may be seen as a snub.
In the acknowledgements of your thesis or dissertation, you should first thank those who helped you academically or professionally, such as your supervisor, funders, and other academics.
Then you can include personal thanks to friends, family members, or anyone else who supported you during the process.
The acknowledgements are generally included at the very beginning of your thesis or dissertation, directly after the title page and before the abstract .
In a thesis or dissertation, the acknowledgements should usually be no longer than one page. There is no minimum length.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the ‘Cite this Scribbr article’ button to automatically add the citation to our free Reference Generator.
George, T. (2022, November 04). Thesis & Dissertation Acknowledgements | Tips & Examples. Scribbr. Retrieved 24 June 2024, from https://www.scribbr.co.uk/thesis-dissertation/acknowledgements/
Is this article helpful?
Tegan George
Other students also liked, dissertation title page, how to write an abstract | steps & examples, dissertation table of contents in word | instructions & examples.
Now Receiving Submissions for CBTS General Research Request for Proposals!
Proposal deadline is Friday, July 26, 2024 at 10 PM CST.
- Skip to primary navigation
- Skip to main content

Cross-Border Threat Screening and Supply Chain Defense
A Department of Homeland Security Center of Excellence

Call for White Papers and Workshop Participation: Supply Chain Resilience Research Needs for Homeland Security
The Center for Accelerating Operational Efficiency (CAOE) at Arizona State University, a Department of Homeland Security (DHS) Center of Excellence, will host a Workshop entitled SCRIPS: Supply Chain Resilience Issues, Problems and Solutions for the Homeland Security Enterprise in Washington D.C. October 1-2, 2024. SCRIPS will bring together leaders from academia, government and industry to identify research needs to ensure the resiliency of critical supply chains. In particular, the workshop will focus on three areas: Ag/Food industry ; Semiconductor Manufacturing ; and Maritime/Port Operations . In preparation for that event, CAOE is soliciting the research community for white papers covering innovative supply chain resilience research ideas and solution methods as they relate to Homeland Security.
Individuals and research groups with relevant interest and expertise are invited to provide white papers as described below. A limited number of respondents will be invited to attend the workshop to further develop prioritized research problem statements and solution approaches. The workshop’s results are anticipated to be used in the development of a Request For Research Proposals (RFP) to be distributed in Spring 2025. Travel expenses will be provided for a limited number of invited participants from accredited academic institutions in the United States.
Submissions will be evaluated based on demonstrated expertise, importance of the problem addressed and the scientific merit and applicability of the solution methodology proposed. Research agendas should have a maximum three-year horizon. Selected contributors will be invited to participate in the October 1-2 workshop.
White Paper Requirements and Acknowledgements:
- White Papers are limited to at most two (2) pages in at least 11 pt font;
- White Paper format should follow the Research Needs/Models/Tools/Solution Template;
- White Papers must be submitted by 11:59pm EDT July 12, 2024 here .
- Submissions must be available for distribution to workshop attendees, publication on workshop website, use in final workshop report and summarization in subsequent RFP. Submission implies such consent.
- Workshop attendance will be by invitation only. Invited participants will be notified by July 31, 2024.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 18 June 2024
Plasma proteomics identify biomarkers predicting Parkinson’s disease up to 7 years before symptom onset
- Jenny Hällqvist ORCID: orcid.org/0000-0001-6709-3211 1 , 2 na1 ,
- Michael Bartl ORCID: orcid.org/0000-0002-7752-2443 3 , 4 na1 ,
- Mohammed Dakna 3 ,
- Sebastian Schade ORCID: orcid.org/0000-0002-6316-6804 5 ,
- Paolo Garagnani ORCID: orcid.org/0000-0002-4161-3626 6 ,
- Maria-Giulia Bacalini 7 ,
- Chiara Pirazzini 6 ,
- Kailash Bhatia ORCID: orcid.org/0000-0001-8185-286X 8 ,
- Sebastian Schreglmann ORCID: orcid.org/0000-0002-4129-5808 8 ,
- Mary Xylaki ORCID: orcid.org/0000-0002-7892-8621 3 ,
- Sandrina Weber 3 ,
- Marielle Ernst 9 ,
- Maria-Lucia Muntean 5 ,
- Friederike Sixel-Döring 5 , 10 ,
- Claudio Franceschi ORCID: orcid.org/0000-0001-9841-6386 6 ,
- Ivan Doykov 1 ,
- Justyna Śpiewak 1 ,
- Héloїse Vinette ORCID: orcid.org/0009-0000-4360-1293 1 , 11 ,
- Claudia Trenkwalder 5 , 12 ,
- Wendy E. Heywood ORCID: orcid.org/0000-0003-2106-8760 1 ,
- Kevin Mills 2 na2 &
- Brit Mollenhauer ORCID: orcid.org/0000-0001-8437-3645 3 , 5 na2
Nature Communications volume 15 , Article number: 4759 ( 2024 ) Cite this article
28k Accesses
2173 Altmetric
Metrics details
- Parkinson's disease
Parkinson’s disease is increasingly prevalent. It progresses from the pre-motor stage (characterised by non-motor symptoms like REM sleep behaviour disorder), to the disabling motor stage. We need objective biomarkers for early/pre-motor disease stages to be able to intervene and slow the underlying neurodegenerative process. Here, we validate a targeted multiplexed mass spectrometry assay for blood samples from recently diagnosed motor Parkinson’s patients ( n = 99), pre-motor individuals with isolated REM sleep behaviour disorder (two cohorts: n = 18 and n = 54 longitudinally), and healthy controls ( n = 36). Our machine-learning model accurately identifies all Parkinson patients and classifies 79% of the pre-motor individuals up to 7 years before motor onset by analysing the expression of eight proteins—Granulin precursor, Mannan-binding-lectin-serine-peptidase-2, Endoplasmatic-reticulum-chaperone-BiP, Prostaglaindin-H2-D-isomaerase, Interceullular-adhesion-molecule-1, Complement C3, Dickkopf-WNT-signalling pathway-inhibitor-3, and Plasma-protease-C1-inhibitor. Many of these biomarkers correlate with symptom severity. This specific blood panel indicates molecular events in early stages and could help identify at-risk participants for clinical trials aimed at slowing/preventing motor Parkinson’s disease.
Similar content being viewed by others

Meta-analysis of shotgun sequencing of gut microbiota in Parkinson’s disease

The power and pitfalls of AlphaFold2 for structure prediction beyond rigid globular proteins
Target engagement and immunogenicity of an active immunotherapeutic targeting pathological α-synuclein: a phase 1 placebo-controlled trial
Introduction.
Parkinson’s disease (PD) is a complex and increasingly prevalent neurodegenerative disease of the central nervous system (CNS). It is clinically characterised by progressive motor and non-motor symptoms that are caused by α-synuclein aggregation predominantly in dopaminergic cells, which leads to Lewy body (LB) formation 1 . The failure of neuroprotective strategies in preventing disease progression is due, in part, to the clinical heterogeneity of the disease—it has several phenotypes—and to the lack of objective biomarker readouts 2 . To facilitate the approval of neuroprotective strategies, governing agencies and pharmaceutical companies need regulatory pathways that use objectively measurable markers—potential therapeutical targets as well as state and rate biomarkers—directly associated with PD pathophysiology and clinical phenotypes 3 .
The recently emerged α-synuclein seed amplification assays (SAA) can identify α-synuclein pathology in vivo and support stratification purposes but still rely on cerebrospinal fluid (CSF) obtained through relatively invasive lumbar punctures 4 . Therefore, this test remains specialised and not readily suitable for large-scale clinical use. As peripheral fluid biomarkers are less invasive and easier to obtain, they could be used in repeated and long-term monitoring, which is necessary for population-based screenings for upcoming neuroprotective trials. While the only emerged serum biomarker in the last years, axonal marker neurofilament light chain (NfL), increases longitudinally and correlates with motor and cognitive PD progression 5 , it is non-specific to the disease process.
Growing data support evidence of PD pathology in the peripheral system, which increases the likelihood of finding a source of matrices for less invasive biomarkers. We know α-synuclein aggregation induces neurodegeneration, which is propagated throughout the CNS. Evidence indicates that additional inflammatory events are an early and potentially initial step in a pathophysiological cascade leading to downstream α-synuclein aggregation that activates the immune system 6 . Inflammatory risk factors in circulating blood (i.e. C-reactive-protein and Interleukin-6 and α-synuclein-specific T-cells) are associated with motor deterioration and cognitive decline in PD 7 , 8 . These inflammatory blood markers can even be identified in plasma/serum samples of individuals with isolated REM sleep behaviour disorder (iRBD), the early stage of a neuronal synuclein disease (NSD), and the most specific predictor for PD and dementia with Lewy bodies (DLB) 6 . NSD was recently proposed as a biologically defined term, for a spectrum of clinical syndromes, including iRBD, PD and DLB, that follow an integrated clinical staging system of progressing neuronal α-synuclein pathology (NSD-ISS) 9 .
In this study, we used mass spectrometry-based proteomic phenotyping to identify a panel of blood biomarkers in early PD. In the initial discovery stage, we analysed samples from a well-characterised cohort of de novo PD patients and healthy controls (HC) who had been subjected to rigorous collection protocols 10 . Using unbiased state-of-the-art mass spectrometry, we identified putatively involved proteins, suggesting an early inflammatory profile in plasma. We thereafter moved on to the validation phase by creating a high-throughput and targeted proteomic assay that was applied to samples from an independent replication cohort, consisting of de novo PD, HC and iRBD patients. Finally, after refining the targeted proteomic panel to include a multiplex of only the biomarkers which were reliably measured, an independent analysis was performed on a larger and independent cohort of longitudinal, high-risk subjects who had been confirmed as iRBD by state-of-the-art video-recorded polysomnography (vPSG), including follow-up sampling of up to 7 years.
In summary, using a panel of eight blood biomarkers identified in a machine-learning approach, we were able to differentiate between PD and HC with a specificity of 100%, and to identify 79% of the iRBD subjects, up to 7 years before the development of either DLB or motor PD (NSD stage 3). Our identified panel of biomarkers significantly advances NSD research by providing potential screening and detection markers for use in the earliest stages of NSD for subject identification/stratification for the upcoming prevention trials.
Proteomic discovery phase 0
We performed a bottom-up proteomics analysis of plasma, which had been depleted of the major blood proteins, using two-dimensional in-line liquid chromatography fractionation into ten fractions and label-free mass spectrometric analysis by QTOF MS E . The discovery cohort consisted of ten randomly selected drug-naïve patients with PD and ten matched HC from the de novo Parkinson’s disease (DeNoPa 10 ) cohort (details can be found in Supplementary Table 1 ). This analysis identified 1238 proteins when restricting identification to originate from at least one peptide per protein and at least two fragments per peptide. After excluding proteins with less than two unique peptides or with an identification score below a set threshold (see method section below), 895 distinct proteins remained. Of these proteins, 47 were differentially expressed between the de novo PD and control groups on a nominal significance level of 95%. Pathway analysis suggested enrichment in several inflammatory pathways. Workflow and Results are shown in Fig. 1 , and 2 Supplementary Figs. 1 , 2.

The study included three phases. Phase 0 consisted of discovery proteomics by untargeted mass spectrometry to identify putative biomarkers, followed by phase I in which targets from the discovery phase were transferred to a targeted, mass spectrometric MRM method and applied to a new and larger cohort of samples, and finally phase II in which the targeted MRM method was refined and a larger number of samples were analysed to evaluate the clinical feasibility of the targeted protein panel.
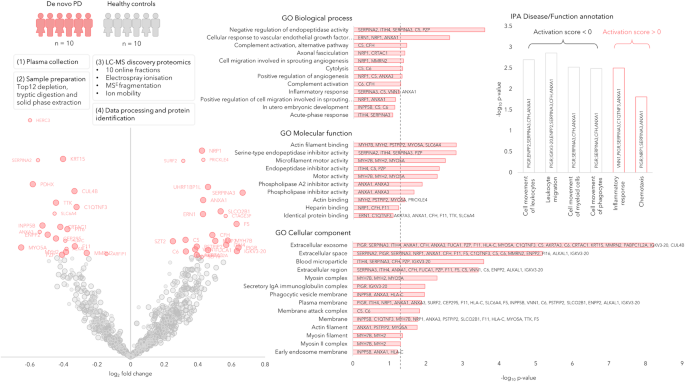
The circle radii in the Volcano plot represent the identification certainty, where large radii represent proteins identified by at least two unique peptides and an identification score >15, smaller radii are given for proteins identified by two or more unique peptides or a confidence score >15. The horizontal axis shows log 2 of the average fold-change and the vertical axis shows −log 10 of the p values. The significantly different proteins are annotated by gene name and coloured in pink, while the non-significant proteins are coloured in grey. GO annotations for the significant proteins are shown, the dashed line represents p = 0.05. Disease and function annotations from IPA are shown, divided into annotations with a positive or negative activation score. Source data are provided as a Source Data file.
Selection of proteins for the targeted proteomic assay
We next developed a validatory, high-throughput and multiplexed, mass spectrometric targeted proteomic assay based on the potential biomarkers identified in the discovery phase. Additional proteins were also included in the assay, several of which had been identified in previous discovery studies of PD, Alzheimer’s disease (AD), and ageing 11 . In addition, we also included several known pro- and anti-inflammatory proteins identified in the literature 12 , 13 , 14 , 15 , which had been previously developed into an in-house targeted proteomic neuroinflammatory panel. Using this approach, we created a targeted proteomic panel, including biomarkers from current scientific developments and preliminary findings from our own work 16 , 17 . This targeted proteomic and multiplexed assay included 121 proteins and aimed to validate biomarkers and probe the pathways identified as being perturbed in the discovery phase. Details can be found in Supplementary Table 2 and Fig. 3 .
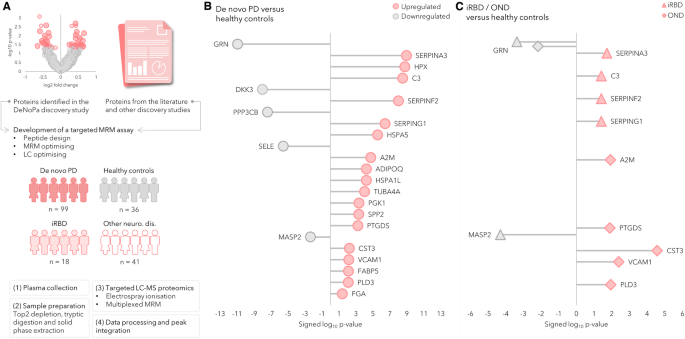
Workflow and overview of the results of the targeted proteomic analysis of de novo Parkinson’s disease (PD) subjects, healthy controls (HC), and the validation cohorts of other neurological disorders (OND) and isolated REM sleep behaviour disorder (iRBD). A A targeted mass spectrometric proteomic assay was developed and optimised. The assay was then applied to plasma samples from cohorts comprising de novo PD ( n = 99) and HC ( n = 36), and validated in patients with OND ( n = 41) and prodromal subjects with iRBD ( n = 18). The protein expression difference between the groups was compared using Mann–Whitney’s two-sided U -test with Benjamini–Hochberg FDR adjustment at 5%. The lollipop charts show the log 10 p values, signed according to fold-changes. Pink icons represent a protein upregulated in an affected group and grey represents a protein upregulated in controls. B Significantly differentially expressed proteins in the comparison between de novo PD and healthy controls. C Significantly differentially expressed proteins between iRBD, OND and HC. Source data are provided as a Source Data file.
Demographics-targeted proteomic validation phase (phase I)
For the targeted proteomics analysis, we used plasma samples, independent from the proteomic discovery step, from 99 individuals recently diagnosed with de novo PD (48 men, 50%, mean age 67 years) and 36 healthy controls (HC; 20 men, 57%, mean age 64 years). This was the main cohort, to which we added further samples for validation that consisted of a heterogeneous group of 41 patients with other neurological diseases (OND) (29 men, 71%, mean age 70 years) and 18 patients with vPSG-confirmed iRBD (10 men, 56%, mean age 67 years). Further details can be found in Table 1 and Fig. 3 .
The identification of biomarkers that were significantly and differentially expressed biomarkers between patients with de novo Parkinson’s disease and healthy controls- Targeted proteomic validation phase (phase I)
Our targeted proteomic assay was developed for 121 proteins, 32 of which we consistently and reliably detected in plasma. Of these 32 markers, 23 were confirmed as being significantly and differentially expressed between PD and HC. We identified six differentially expressed proteins in the comparison between iRBD patients and HC and between OND and HC (Fig. 3 ). Both the de novo PD and iRBD groups demonstrated an upregulated expression of the serine protease inhibitors SERPINA3, SERPINF2 and SERPING1, and of the central complement protein C3. Granulin precursor protein was shown to be downregulated in all three patient groups (PD, iRBD and OND) compared to HC. The OND and PD groups had a shared and upregulated expression of the proteins PTGDS, CST3, VCAM1 and PLD3. Detailed information about the diagnoses of the OND group can be found in Table 1 , and detailed information about the proteins can be found in Supplementary Table 2 . Figure 4 shows the significantly different proteins as Box-scatter plots.

The data are displayed as Box and Whisker plots overlaid with scatter plots of the individual measurements. The whiskers show the minimum and maximum, and the boxes show the 25th percentile, the median and the 75th percentile. The protein expression difference between the groups was compared using Mann–Whitney’s two-sided U -test with Benjamini–Hochberg multiple testing correction (FDR adjustment at 5%). ns not significant, * p < 0.05, ** p < 0.01, *** p < 0.001 and **** p < 0.0001. The proteins are represented by gene names. Source data are provided as a Source Data file.
The biological significance of the differentially expressed proteins- Targeted proteomic validation phase (phase I)
The involvement of the differentially expressed proteins and their impact on biological processes were evaluated using pathway analysis (Ingenuity Pathway Analysis [IPA], Qiagen). The significantly differentially expressed proteins between PD and HC were used as input, with a fold-change set as the expression observation. We considered pathways as significant if they had an enrichment p value <0.05. At least two of the input proteins were included. Three major pathway clusters were identified and consisted of (i) the expression of serine protease inhibitors or serpins and complement and coagulation components, (ii) endoplasmic reticulum (ER) stress/heat shock-related proteins and (iii) the expression of VCAM1, SELE and PPP3CB. The highest enrichment scores were identified in the pathways acute phase response signalling ( p = 7.8 E −10 ), coagulation system ( p = 7.4 E −6 ), complement system ( p = 8.1 E −6 ), LXR/RXR activation ( p = 9.1 E −6 ), FXR/RXR activation ( p = 9.8 E −6 ) and glucocorticoid receptor signalling ( p = 2.0 E −5 ). These are all pathways involved in inflammatory responses. We also identified pathways related to the unfolded protein response ( p = 0.004) and neuroinflammation ( p = 0.04), although with lower enrichment scores. For details, see Supplementary Fig. 1 .
Inflammation-related pathways (including both the complement system and the acute phase response) demonstrated the highest significance levels, followed by pathways regulating protein folding, ER stress, and heat shock proteins. A network representation of proteins and pathways showed clusters consisting of inflammation/coagulation/lipid metabolism (FXR/RXR and LXR/RXR), heat shock proteins/protein misfolding, and more heterogenous pathway clusters related to Wnt-signalling and extracellular matrix proteins. Figure 5 illustrates the potential detrimental and protective mechanisms suggested to be taking place based on the protein expressions observed in this study, leading to oligomerisation and accumulation of α-synuclein in neuronal Lewy body inclusions and, finally, dopaminergic neuronal cell loss.

Oligomerisation and accumulation of α-synuclein in Lewy body inclusions is a key process in the pathophysiology of neuronal synuclein disease, i.e. Parkinson’s disease and dementia with Lewy bodies from aggregation and accumulation, the pathological pathway includes different steps finally leading to the loss of dopaminergic neurons. Protective and detrimental mechanisms influence these processes, based on the differently expressed protein profiles, assessed by targeted mass spectrometry in our study. Detailed information about the proteins can be found in Supplementary Table 2 .
Multivariate analysis shows differences between the proteomes of Parkinson’s disease and controls- Targeted proteomic validation phase (phase I)
Principal component analysis (PCA) demonstrated that the HC and PD groups formed two clusters separate from each other over the first and second principal components (PC), attributed with 23.5% and 13.9% of the model’s total variance, respectively. The iRBD group was situated in the middle of HC and PD, and the OND group varied considerably with no evident clustering, as expected due to the heterogeneity of diseases. The corresponding loadings of PC1 and PC2 demonstrated that those with PD correlated with lower levels of PPP3CB, DKK3, SELE and GRN, and higher levels of most of the other proteins. The loadings plot had a high level of covariation in the expression of the PPP3CB, DKK3 and SELE proteins, which were all downregulated in PD. These proteins correlated negatively with the expression of SERPINs, complement C3 and HPX, which all showed a high degree of covariation, and were upregulated in the PD group. Data are displayed in Supplementary Fig. 2 .
The use of multiplexed protein panels of protein biomarkers for the prediction of de novo Parkinson’s disease- Targeted proteomic validation phase (phase I)
We next applied machine learning to construct a discriminant OPLS-DA model using the PD and HC samples from the validation phase. The samples clustered into two distinct and well-separated classes, and evaluation of the model showed that it was highly significant ( p = 2.3E −27 permutations p = <0.001). The proteins with the greatest influence on the class separations were GRN, DKK3, C3, SERPINA3, HPX, SERPINF2, CAPN2, SERPING1 and SELE. We predicted the iRBD samples in the model, which resulted in 13 subjects classified as PD (72%) and five not belonging to either group. None of the iRBD samples were classified as controls. We additionally predicted the OND samples, out of which nine were classified as HC, 12 as PD and 19 were not classified as belonging to either group. The 12 samples predicted as PD did not demonstrate enrichment according to the OND groups. The random distribution of the OND samples between PD and HC indicates that the heterogenous group of OND individuals does not share a distinct protein expression with either the HC or PD groups. The iRBD samples that were classified as PD, and not as HC, strongly suggest a shared proteomic profile between iRBD and the protein expression observed in the newly diagnosed PD patients.
We subsequently explored if the observed protein expressions could be used to build a regression model capable of predicting whether individuals belonged to the PD or HC groups. We identified a panel of proteins that discriminated between PD and HC with 100% accuracy and then constructed a linear support vector classification model and applied recursive feature elimination to pinpoint the most discriminating variables. The data were divided into two parts: one consisting of 70% for model training and one containing 30% for testing. The proportion of PD and control samples was maintained in each part. The number of features included in the model was determined by feature ranking with cross-validated recursive feature elimination in the training dataset. The feature selection resulted in a model with eight predictors: GRN, MASP2, HSPA5, PTGDS, ICAM1, C3, DKK3 and SERPING1. The training data were predicted in the model and resulted in all samples being classified in the correct class. We further constructed receiver operating characteristic (ROC) and precision-recall (PR) curves to illustrate the ability of each protein to distinguish between PD and HC and compared this with the ability of the combined multiplexed protein panel. The combined panel achieved an AUC of 1.0 on both ROC and PR curves. The AUC of the individual predictors ranged from 0.53 to 0.92 in the ROC curve, and from 0.79 to 0.96 in the PR curve (Fig. 6 ). We further evaluated the whole dataset by performing repeated cross-validation with six splits of the data and 40 repetitions. The resulting classification metrics (Supplementary Fig. 3 ) demonstrated average and standard deviation for precision, recall, F1 score, and balanced accuracy score of 0.87 ± 0.09, 0.87 ± 0.08, 0.86 ± 0.09 and 0.82 ± 0.12, respectively, thereby indicating a highly robust classification model. Testing the model’s specificity for PD, we predicted the heterogenous group of OND, resulting in 26 of the 42 samples being classified as PD-like. Prediction of the prodromal iRBD group resulted in 17 of 18 samples being classified as PD-like. We compared the prediction of the OND and iRBD samples between the OPLS-DA and SVM models, finding that most of the samples were classified in the same group in both models (out of the samples with a classification in the OPLS-DA model: 82% in OND and 100% in iRBD). The proportion of iRBD samples classified as PD in our models (72% in the OPLS-DA model and 94% in the SVM model) is in line with clinical evidence based on longitudinal cohort studies, reporting that over 80% of iRBD subjects will develop an advanced NSD with motor impairment and/or cognitive decline 18 . We evaluated the influence of age and sex on the proteins included in the support vector model and found that neither influenced the model’s classification ability (see Supplementary Methods 2 for details).

The model was trained on 70% of the samples to establish the most discriminating features. Applying cross-validated recursive feature elimination, the top predictors were determined as a granulin precursor, mannan-binding lectin-serine peptidase 2, endoplasmic reticulum chaperone-BiP, prostaglandin-H2 d -isomerase, intercellular adhesion molecule-1, complement C3, dickkopf-3 and plasma protease C1 inhibitor. The remaining 30% of samples were predicted in the model and resulted in 100% prediction accuracy. Receiver operating characteristics (ROC) and precision-recall (PR) curves of the individual and combined proteins in the test set demonstrated that the individual proteins achieved ROC area under the curve (AUC) values 0.53–0.92 and PR values 0.79–0.96, while the combined predictors reached an area under the curve = 1.0. Source data are provided as a Source Data file.
Development of a rapid and refined LC-MS/MS method and evaluation of an independent and longitudinal iRBD cohort (Independent replication cohort-phase II)
To evaluate the results from the initial prediction models focusing on at-risk subjects, we developed and refined our targeted and multiplexed proteomic test to quantitate only those proteins that were readily and reliably detectable from the initial targeted proteomic assay ( n = 32). Next, we analysed an additional set of 146 longitudinal samples from an independent cohort of 54 individuals with iRBD. This cohort was available from continuing recruitment at the same centre and consisted of longitudinally followed iRBD subjects. Deep phenotyping revealed 100% (54/54) had RBD on PSG, 88.9% (48/54) had hyposmia as identified with the Sniffin’ Stick Identification Test, and 91.7 % (22/24) had neuronal α-synuclein positivity as shown by α-synuclein Seed Amplification Assay (SAA) in cerebrospinal fluid (CSF) 19 . Longitudinal follow-up was available for up to 10 years, during which 16 subjects (20%) phenoconverted to either PD ( n = 11) or dementia with Lewy bodies (DLB; n = 5). Since only serum samples were available from the independent replication cohort (further details can be found in Supplementary Table 3 ), we investigated how the proteins in our assay correlated between plasma, serum, and CSF and found good correlations between plasma and serum, but poor correlations between these blood matrices and CSF. The limited correlations between blood and CSF proteins correspond to those of other studies comparing the protein expression between plasma/serum and CSF 20 , 21 and underscore that our test does not necessarily reflect a prodromal and PD-specific proteomic signature of the protein expression in the CSF in proximity to the brain, but rather shows an earlier change in the blood protein expression between healthy status and very early PD patients (Details from this comparison can be found in Supplementary Methods 1 and Supplementary Fig. 4 ).
We applied all available longitudinal iRBD samples ( n = 146) from phase II to the two machine-learning models (OPLS-DA and support vector machine) constructed in phase I (PD vs. HC). The OPLS-DA model, based on all 32 detected proteins, identified 70% of the iRBD samples as PD, while the SVM model, which was based on a panel of eight proteins, identified 79% of the samples as PD. As mentioned above, at the time of analysis, 16 of the 54 subjects in our longitudinal iRBD validation cohort had developed PD/DLB. The earliest correct classification was 7.3 years prior to phenoconversion and the latest was 0.9 years prior to diagnosis (average 3.5 ± 2.4 years). Detailed information can be found in Fig. 7 and Supplementary Methods 3 .
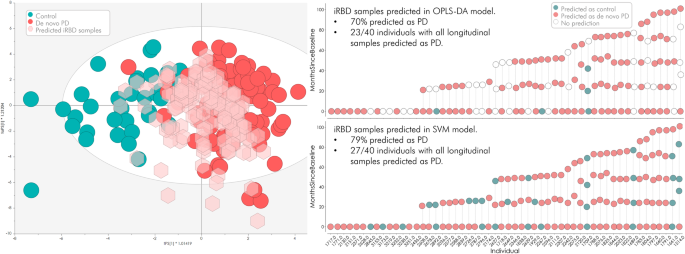
146 new serum samples from individuals diagnosed with iRBD, several with longitudinal follow-up samples, were predicted in the OPLS-DA model. 70% of the samples were predicted as Parkinson’s disease (PD), and 23 of 40 individuals had all their longitudinal samples predicted as PD. In the more refined support vector machine (SVM) model, 79% of the 146 new samples were predicted as PD and 27 of 40 individuals consistently had all their longitudinal samples predicted as PD. Source data are provided as a Source Data file.
The correlation between differentially expressed protein biomarkers and patients’ clinical data in the targeted proteomic validation phase (phase I)
We next evaluated the relationship between proteins and clinical data by correlating the protein expression in PD and HC (from phase I) with clinical scores (Mini-Mental State Examination [MMSE], Hoehn & Yahr stage [H&Y] and UPDRS [Unified Parkinson’s Disease Rating Scale; I–III, and total score]). We found negative correlations for GRN, DKK3, PPP3CB, and SELE with H&Y and UPDRS parts II, III, and total score, possibly indicating a connection between a more severe clinical (especially motor) impairment and lower expression of markers in the Wnt-signalling pathways (DKK3 and PPP3CB). Higher Cystatin C plasma levels correlated with higher numbers in UPDRS part III (motor performance) and UPDRS total score. The same was found for PTGDS plasma levels, which were also negatively correlated with MMSE. The central complement cascade protein, C3, negatively correlated with MMSE, and positively correlated with H&Y, UPDRS part III, and total score. The UPR-regulating protein BiP (HSPA5) correlated negatively with MMSE, and positively with H&Y and UPDRS parts II, III, and total score. The ERAD-associated proteins, HSPAIL and adiponectin, were positively correlated with H&Y, and UPDRS parts II, III, and total score. SERPINs (SERPINA3, SERPINF2 and SERPING1) and hemopexin (HPX) correlated negatively with MMSE and positively with H&Y and UPDRS parts II, III, and total score. In general, the MMSE score was inversely correlated with H&Y stage and UPDRS scores. For detailed information, see Fig. 8 and Table 2 .

The correlation was performed using Spearman’s procedure, and the clustering method was set to average. The clustering metric was Euclidean. The heatmap is coloured by correlation coefficient where red represents positive and blue negative correlations. The proteins are represented by gene names. Detailed information about the protein correlations can be found in Supplementary Table 3 . De novo Parkinson’s disease ( n = 99) and healthy controls ( n = 36). MMSE mini-mental state examination, UPDRS unified Parkinson’s disease rating Scale. Source data are provided as a Source Data file.
Comparison of clinical outcomes and measurements in the longitudinal iRBD cohort-Independent replication cohort-phase II
The longitudinal expression in the iRBD samples was evaluated using linear mixed-effects models. Conditional growth models with random slopes and random intercepts between the individuals were constructed. After adjusting the p values for multiple testing by applying the Benjamini–Hochberg (BH) procedure with alpha = 0.05, we found that Butyrylcholinesterase (BCHE) was significantly decreased over the timepoints in the iRBD individuals ( p = 0.01). We next focused only on the iRBD samples with at least two timepoints and for which PD had consistently been predicted in the SVM model ( n = 90). This produced comparable results to the initial model with BCHE significantly related with time since baseline ( p = 0.01), but also TUBA4A was nominally significantly increased ( p = 0.04) although not passing the BH FDR threshold. The modelling also demonstrated that the clinical measurements H&Y ( p = 0.02), UPDRS I–III ( p = 0.02), and UPDRS I and III ( p = 0.03 and 0.03, respectively), were significantly related to the time since baseline in the iRBD group post multiple testing correction. PD non-motor symptoms, as measured on the PD NMS sum score, were strongly correlated with longitudinal motor progression ( p = 5E −8 ). Similarly, the questionnaire for quality of life PDQ-39’s mean values also correlated with longitudinal motor progression ( p = 0.005). From available routine blood values, cholesterol was associated with longitudinal timepoints ( p = 0.02). Details can be found in Supplementary Table 4 . Correlating the clinical measurements with the targeted proteomic data, we applied Spearman’s correlation and found that cholesterol was positively correlated with six of the identified proteins (Supplementary Table 5 ), including HSPA8, APOE and MASP2 ( p = 5E −9 , 0.0003 and 0.003, respectively). Also significantly correlated, but to a lesser degree and not passing the BH FDR threshold, were the PD NMS sum which correlated negatively with TUBA4A (p unadjusted = 0.01) and the PDQ-39 mean values, which correlated negatively with CST3 and PTGDS ( p unadjusted = 0.03 and 0.05, respectively).
PD has emerged as the world’s fastest-growing neurodegenerative disorder and currently affects close to 10 million people worldwide. Consequently, there is an urgent need for disease-modifying and prevention strategies 22 , 23 . The development of such strategies is hampered by two limitations: there are major gaps in our understanding of the earliest events in the molecular pathophysiology of PD, and we lack reliable and objective biomarkers and tests in easily accessible bio-fluids. We, therefore, need biomarkers that can identify PD earlier, preferably a significant time before an individual develops significant neuronal loss and disabling motor and/or cognitive disease. Such biomarkers would advance population-based screenings to identify individuals at risk and who could be included in upcoming prevention trials.
In the last years, CSF SAA emerged as the most specific indicator for NSD, in prodromal stages like iRBD, with an impressively high sensitivity and specificity of up to 74 and 93%, respectively, across various cohorts 9 , 24 . Despite the many questions surrounding SAA that need to be answered, including the ultimate understanding of its functionality, it is a true milestone for advancing prevention trials. It is, however, hampered by having only been shown to be robust in CSF and by the slow development and high variability of SAA in peripheral blood 25 , as well as by the lack of quantification capabilities. An easier and more accessible biofluid test would enable screening large population-based cohorts for at-risk status to develop an NSD. Therefore, the identification of additional biomarkers is needed, as is further knowledge of the biomarkers and pathways of the underlying pathophysiology (e.g. inflammation) during the earliest stage of NSD.
Other emerging multiplex technologies are increasingly used to identify individual proteomic biomarkers. However, these techniques are not true proteomic or ‘eyes open’ methods, as they rely on selected large panels of specific antibodies/and other (e.g. aptamer)-based assay technologies. These techniques, although useful, have not provided consistent results 3 , 26 . Proteomics using mass spectrometry measures all expressed proteins in an unbiased fashion as opposed to those selectively included in a panel that also includes variability due to cross-reactivity. Therefore, proteomic screening using mass spectrometry-based techniques is much more likely to identify pathways or biomarkers and provides more meaningful insights into the disease mechanisms involved in PD. We found a discrepancy between the detected markers during the discovery and the targeted phases. This is a known phenomenon in biomarker translation 27 that is also reflected in the low number of biomarkers having received FDA approval 28 . We addressed this by using previously reported successful improvement strategies in proteomic approaches, namely by refining our panel, reducing the number of markers, and increasing the sample size 29 . Furthermore, the validation of potential biomarkers was performed on a second and different type of mass spectrometer (triple quadrupole), which has the advantage of being available in all large hospitals.
Targeted MS has been previously applied in PD, including by the current authors, but the biological fluid used in the majority of studies is CSF 30 and not peripheral fluids such as blood. Here we demonstrate that even with a very low required volume of plasma/serum (10 µl) targeted proteomic is feasible.
The targeted proteomic assay presented here was developed from proteins identified in an unbiased discovery study, from our previous research, and from the literature. It included several inflammatory markers, Wnt-signalling members, and proteins indicative of protein misfolding. When analysing PD, OND, iRBD and HC in the targeted proteomic validation phase, we identified and confirmed 23 distinct and differentially expressed proteins between PD and HC. Our analysis moreover demonstrated that iRBD possesses a significantly different protein profile compared to HC, consisting of decreased levels of GRN and MASP2 and increased levels of the complement factor C3 and SERPINs (SERPINA3, SERPINF2 and SERPING1), thus indicating early involvement of inflammatory pathways in the initial pathophysiological steps of PD. Comparing these results to previous findings by our and other groups 8 , 31 highlights the link between these proteins and the pathways of complement activation, coagulation cascades, and Wnt-signalling.
By applying machine-learning models, we classified and separated de novo PD or control samples with 100% accuracy based on the expression of eight proteins (GRN, MASP2, HSPA5, PTGDS, ICAM1, C3, DKK3 and SERPING1).
With an independent validation, we added (a) a larger sample set and (b) longitudinal samples from the most interesting subgroup with 54 iRBD subjects and a total of 146 serum samples. We were able to validate our previous panel with a high prediction rate (79%) of these individuals as seen in PD in the targeted approach. Interestingly, the biomarker panel itself did not correlate with longitudinal expression but remained robust after the initial classification of iRBD. So far, 16 of the 54 iRBD subjects converted to PD/DLB (stage 3 NSD). Out of these samples, the SVM model predicted ten individuals with all their timepoints classified as PD, and of the 11 iRBD subjects who converted to PD/DLB, eight were identified as PD by the proteome analysis. Our panel, therefore, identified a PD-specific change in blood up to 7 years before the development of the stage 3 NSD.
The main shortcoming with many previously explored PD biomarkers is weak or no correlation with clinical progression data. So far, outcome measures in clinical trials are primarily based on motor progression, often by a clinical rating scale such as the UPDRS and/or wearable technologies. More objective biomarkers correlating with or reflecting the progression of the pathophysiology and clinical symptoms would be of the utmost importance. We, therefore, calculated correlations with clinical parameters and identified an association with multiple markers, including DKK3, PPP3CB and C3, indicating downregulation of Wnt-signalling pathways. Increased activity of the complement cascade correlated with higher scores in symptom severity (UPDRS part III and total score) and lower scores in cognitive performance (MMSE).
Protein (i.e. α-synuclein) misfolding is a well-known component of PD pathology and is believed to be the key factor behind Lewy body formation 32 . The transport of excessive amounts of misfolded proteins or increased folding cycles can induce ER stress. A cellular defence mechanism to alleviate ER stress is the unfolded protein response (UPR) reducing ER protein influx and increasing protein folding capacity 33 . The UPR is mainly activated by BiP-bound misfolded proteins 34 . The higher expressed markers HSPA5 (UPR-regulating protein BiP) and HSPA1L in our plasma samples of early PD indicate ER stress as a significant factor in the disease process and has been previously linked to PD in both mouse models and brain tissue studies 35 , 36 .
As mentioned by other groups and confirmed in our results, increasing evidence suggests inflammation is a specific feature in early PD. Complement activation has been associated with the formation of α-synuclein and Lewy bodies in PD and deposits of the complement factors iC3b and C9 have been found in Lewy bodies 37 . C3 is a central molecule in the complement cascade and was highly upregulated in blood in both PD and both independent iRBD sample sets analysed in this study. This upregulation in the earliest phase of motor PD (stage 3 NSD), and even in the prodromal phase (stage 2 NSD), clearly indicates inflammation as an early, if not the initial, event in PD neurodegeneration. Complement C3 levels correlated positively with indicators of motor dysfunction (H&Y stage and UPDRS)—indicating a direct connection between high plasma levels of inflammatory proteins and motor symptoms—and negatively with cognitive decline, here with the MMSE.
The protein Mannan-binding serine peptidase 2 (MASP2), an initiator of the lectin part of the complement cascade, was significantly downregulated in PD and iRBD. MASP1 and MASP2 proteins are inhibited by plasma protease C1 inhibitor SERPING1 in the lectin pathway, with SERPING1 modulating the complement cascade as it belongs to the SERPIN family of acute phase proteins 38 . In experimental PD mice models, increased SERPING1 levels are associated with dopaminergic cell death 39 . Acting as a serine/cysteine proteinase inhibitor, SERPING1 can increase serine levels, which could also affect αSyn phosphorylation. This can play a crucial role in PD pathology, as almost 90% of αSyn in Lewy bodies is phosphorylated on Serine129 40 , 41 . We identified increased SERPING1 plasma levels in both PD and iRBD in our analysis (compared to HC), thus contributing to conditions with increased αSyn phosphorylation, consecutive aggregation, Lewy body formation, and finally degeneration of dopaminergic neurons. Furthermore, we observed a strong correlation of SERPING1 plasma levels with UPDRS II, III and total score, as a direct measure of dopaminergic cell loss 39 .
Alpha-2-antiplasmin (SERPINF2) was also significantly upregulated in PD and iRBD. SERPINF2 is a major regulator of the clotting pathway, acting as an inhibitor of plasmin, a serine protease formed upon the proteolytic cleavage of its precursor, plasminogen, by tissue-type plasminogen activator (t-PA) or by the urokinase-type plasminogen activator (u-PA). Plasmin has been reported to cleave and degrade extracellular and aggregated αSyn 42 . Recently, we showed that activation of the plasminogen/plasmin system is decreased in PD, indicated by decreased plasma levels of uPA and its corresponding receptor uPAR, while t-PA was associated with faster disease progression 8 . The upregulation of SERPINF2 observed here is another indicator of decreased plasmin activity. Alpha-1-antichymotrypsin (SERPINA3), a third member of the SERPIN family, was also upregulated in the PD subjects. In the CNS, the primary source of SERPINA3 is astrocytes, where its expression is upregulated by various inflammatory receptor complexes 38 .
Overall, independent upregulation of these three members of the SERPIN (SERPING1, SERPINF2, SERPINA3) family is also indicative of increased inflammatory activity, combined with less activation of the plasmin system, and correlation with motor and non-motor symptom severity. In addition, a strong downregulation of progranulin ( GRN ) was detected, indicating a potential loss of neuroprotection and increased susceptibility to neuroinflammation. GRN may act as a neurotrophic factor, promoting neuronal survival and modulating lysosomal function. Loss-of-function mutations in the GRN gene are a cause of frontotemporal dementia and familial DLB. GRN gene variants are also known to increase the risk of developing Alzheimer’s disease (AD) and PD 43 . The main characteristics of neurodegeneration related to GRN are TDP43(-Transactive response DNA binding protein 43) inclusions, but Lewy body pathology is also very common. Loss of progranulin has further been linked to increased production of pro-inflammatory species such as tumour necrosis factor (TNF) and IL-6 in microglia 15 . A study in mice showed that Grn -/- mice had elevated levels of complement proteins, including C3, even before the onset of neurodegeneration 44 . Additionally, previous studies have found GRN downregulated in serum samples of advanced PD compared to AD and healthy individuals 45 .
As a possible compensatory reaction to the described increased inflammatory markers, the levels of Prostaglandin-H 2 d -isomerase (PTGDS)/Prostaglandin-D 2 synthase (PGDS2), better known as β-trace protein, were upregulated. PDGDS is an important brain enzyme producing prostaglandin D2 (PGD2), which has a neuroprotective and anti-inflammatory function. The upregulation reported here could be a reaction to the amount of neuronal cell loss, which is also seen in the significant correlation with the clinical motor and cognitive scales (see below). Furthermore, β-trace protein is a marker for CSF and is used to identify the fluid in clinical routine diagnostics, thus helping detect CSF leakage 46 . Increased plasma levels could be indicative of a disrupted blood–brain barrier (BBB), often discussed in PD pathology 47 and demonstrated in our cohorts.
Our study shows that the Wnt-related proteins DKK3 and PPP3CB are strongly downregulated in de novo PD. DKK3 is an activator of the canonical Wnt/β-catenin branch and PPP3CB is a component of the non-canonical Wnt/Ca 2+ signalling pathway. Wnts are secreted, cysteine-rich glycoproteins that act as ligands to locally stimulate receptor-mediated signal transduction of the Wnt-pathway 48 . Wnt-signalling is crucial for the development and maintenance of dopaminergic neurons 49 , shows protective effects on midbrain dopaminergic neurons 50 , and seems to be involved in the maintenance of the BBB 48 , 51 . Wnt-ligands and agonists trigger a “Wnt-On” stage, characterised by neuronal plasticity and protection, while the opposite “Wnt-Off” stage, potentially leading to neurodegeneration, triggered by the phosphorylation activity of glycogen synthetase kinase-3β (GSK-3beta) 50 , 52 . Wnt-inhibitors are separated into secreted Frizzled-related proteins (sFRP) and Dickkopf proteins (DKK). DKK1, DKK2 and DKK4 act as antagonists, while DKK3 is an agonist and activator 53 . Adult neurogenesis is primarily governed by canonical Wnt/β-catenin signaling 54 and downregulation of Wnt-signalling promotes dysfunction and/or death of dopaminergic neurons. Restoration of dopaminergic neurons was shown in mice where β-catenin was activated in situ 52 and neural stem cells transplanted to the substantia nigra of medically PD-induced mice induced re-expression of Wnt1 and repair dopaminergic neurons 55 . DKK3 and PPP3CB were strongly downregulated in de novo PD, removing an important line of defence against the detrimental loss of dopaminergic neurons. The downregulation of the Wnt-signalling pathways was further correlated with higher motor scores (UDPRS and H&Y stages).
Wnt-signalling in PD is not only promising as a potential biomarker. In oncology, drugs can modify Wnt-pathways, which is of interest to the PD field 56 . Some substances show no BBB-permeability. As a disrupted BBB seems to be apparent in PD, these drugs may be effective. Furthermore, these substances are also relevant for PD treatment: research points towards a peripheral starting point of PD and future therapies should be administered as early as possible 57 . These promising substances include DKK- as well as GSK inhibitors, but to date, no drugs targeting the Wnt-signalling pathways have been effectively tested in clinical trials, including in those with neurodegenerative diseases. Progress and clinical trials are urgently needed here.
The transfer of multi-omics analysis to clinically meaningful results that directly impact future drug trial planning and biomarker validation, depends fundamentally on correlating these results and altered pathway regulations with established clinical scores. The markers we analysed in our targeted mass spectrometry panel did not only show different expression patterns between HC, PD, and in both of our independent iRBD sample sets, but most of the markers also robustly correlated with important clinical scores (UPDRS and MMSE, see Table 1 ). Cognitive decline correlated negatively with the SERPINs and complement factor C3. The burden of motor and non-motor symptoms and overall symptom severity rated by UPDRS and its subscores correlated positively with the SERPINs, Complement C3, and negatively with DKK3, GRN, and SELE. So, increased inflammatory activity and downregulation of Wnt-signalling seem to strongly affect the clinical picture of PD subjects.
The iRBD subjects showed decreased levels of BCHE over time compared to controls. BCHE has been reported as decreased in serum samples of PD with cognitive impairment 58 . Validation of this easily assessable marker in serum is needed to evaluate its predictive potential.
While we did not find significant differences when we compared paired serum and plasma samples; the analysis of paired samples of plasma/serum and CSF only correlated weakly with the marker concentrations in these peripheral and central compartments. This discrepancy has been reported by several groups 20 , 21 . One reason is that mass spectrometry-based proteome analysis is always biased towards quantification and detection of the most abundant proteins in each sample matrix, and the total protein concentrations in human plasma/serum are more than two orders of magnitude higher than that in CSF. Further, the regulatory function of the blood–brain barrier seems to play a different role for different proteins, as some, like c-reactive protein, show a strong correlation between CSF and plasma, but most of the proteins do not. CSF and blood proteome show complex dynamics influenced by multiple and still mostly unknown factors. The protein shift in samples with a known BBB dysfunction (determined by the CSF/serum albumin index or the CSF/plasma ratio) can not be determined for individual proteins nor the dysfunction be localised by mass spectrometry 20 .
Our model could not correctly predict phenoconversion in all cases. The reasons for this can be varied: The proteome pattern changes over time and the period between sampling and phenconversion may play a role. The three PD phenoconverters that were not predicted as PD neither differ clinically or demographically from the phenoconverters, nor from the non-phenoconverters. iRBD diagnosis in our study was confirmed by vPSG, supported by a high percentage of additional measurements including hyposmia and CSF SAA positivity. Therefore, even those iRBD cases that do not show the PD-proteome pattern still have a high-risk constellation of converting to PD/DLB on three different levels (PSG, olfaction, and SAA). Continuing further longitudinal follow-up of these subjects will elucidate our understanding of when and potentially why conversion occurs/does not occur. It is known that around 80% of iRBD subjects develop NSD, i.e. PD/DLB, with a rate of 6% per year, as shown in a multicenter cohort including ours 59 . To a lesser extent, iRBD subjects develop the intracytoplasmic glial α-synuclein aggregation disorder Multiple Systems Atrophy (MSA) 59 , 60 . Although RBD is common in MSA (summary prevalence of 73% 61 ), none of our iRBD subjects have, as yet converted to MSA. Recruiting and following large longitudinal at-risk cohorts is, therefore, very important and future studies will not only identify biomarkers for phenoconversion from stage 1 or 2 to eventually stage 3 NSD or MSA, but also identify the many possible factors of resilience (including genetics, etc.) of NON-conversion which will be as, if not more important than identifying indicators for phenoconversion. Both direction progression biomarkers from stage 1 and 2 cohorts will have tremendous implications for future neuroprevention trials as phenoconversion itself is (due to the low annual rate) unlikely to be an outcome measure.
A significant strength of our biomarker discovery to translation pipeline is that it allows for the developed test to be easily validated and translated to any clinical laboratory equipped with a tandem LC-MS instrument. One advantage of using triple quadrupole platforms is that additional and better biomarkers can easily be augmented into the test described in this manuscript. Thus, any test could be refined and optimised over time with very little modification to the assay as additional biomarkers are discovered. Clinical testing for neurological disorders is limited to the use of a selected few well-characterised individual markers and translating biomarkers to eventual clinical application is notoriously challenging. The power of using multiplexed biomarker technologies with machine learning enables biomarkers to be evaluated in context with other markers of pathological events, thereby creating a ‘disease profile’ as opposed to individual markers. This approach opens the biomarker discovery field for many disorders and increases the specificity and sensitivity of testing, as demonstrated in this study. The combination of multiplexed analysis of biomarker panels analysed on triple quadrupole platforms can advance biomarker translation to clinical application; this mass spectral technology is already embedded in many clinical diagnostics labs for routine small molecule analyses.
Our peripheral blood protein pattern for PD helps not only to classify but also to predict the earliest stage of the disease. We find differently expressed proteins in pre-motor iRBD and early motor stages of the disease compared to HC. Multiple markers also correlated with the progression of motor and non-motors symptoms. Thus, our blood panel can also identify subjects at risk (stage 2) to develop PD up to 7 years before advancing to motor stage 3. Next steps will be the independent validation in other (and even earlier) non-motor cohorts, e.g. in subjects with hyposmia also at-risk for PD 62 and in our population-based Healthy Brain Ageing cohort in Kassel 63 . It would further be interesting to evaluate the predictive potential of these identified markers with continuing clinical follow-up and together with other established PD progression markers like serum neurofilament light chain 5 and dopamine transporter imaging in a longitudinal analysis.
Our work was predominantly focused on the similarities between PD and iRBD. The authors are unaware of any study that has analysed longitudinally collected samples and prodromal cohorts, including iRBD and phenoconverters. Future work would include (i) validation of our findings in independent cohorts consisting of iRBD and other at-risk subjects for the synuclein aggregation disorders in neurons (PD, DLB) and oligodendrocytes (MSA), (ii) refinement of the panels of biomarkers developed in this study including sensitivity and technical performance, (iii) and using the pipeline described in this manuscript, the identification and validation of additional biomarkers that could distinguish between the different clinical syndromes with the ultimate goal of identifying progression biomarkers as outcome measures for prevention trials.
In summary, instead of single biomarkers, in a univariate approach, we have created a pipeline using a targeted proteomic test of a multiplexed panel of proteins, together with machine learning. This powerful combination of multiple well-selected biomarkers with state-of-the-art machine-learning bioinformatics, allowed us to use a panel of eight biomarkers that could distinguish early PD from HC. This biomarker panel provided a distinct signature of protective and detrimental mechanisms, finally triggering oxidative stress and neuroinflammation, leading to α-synuclein aggregation and LB formation. Moreover, this signature was already present in the prodromal non-motor (stage 2 NSD), up to 7 years before the development of motor/cognitive symptoms (stage 3), supporting the high specificity of iRBD and its high conversion rate to PD/DLB 18 . Most importantly, this blood panel can, in the future, upon further validation help identify subjects at risk of developing PD/DLB and stratify them for upcoming prevention trials.
Patient cohorts and sample collection and processing
Our research complies with all relevant ethical regulations. Institutional review board statements were obtained from the University Medical Centre in Goettingen, Germany, Approval No. 9/7/04 and 36/7/02. The study was conducted according to the Declaration of Helsinki, and all participants gave written informed consent. All plasma, serum and CSF samples from subjects were selected from known cohorts using identical sample processing protocols designed by the Movement Disorder Center Paracelsus-Elena-Clinic.
Patients with de novo PD were diagnosed according to the UK Brain Bank Criteria, without PD-specific medication. Diagnosis in all subjects was supported by (1) a positive (i.e. >30% improvement of UPDRS III after 250 mg of levodopa) acute levodopa challenge testing 64 in all PD subjects, (2) hyposmia by smell identification test (Sniffin Sticks 65 ) in all PD subjects and (3) 1.5-tesla Magnetic Resonance Imaging (MRI) without significant abnormalities or evidence for other diseases in all but three subjects who were excluded (due to significant vascular lesions or evidence for hydrocephalus) from the analysis. Participants not fulfilling the above criteria and meeting criteria for other neurological disorders were named as other neurological disorders (OND). OND consists of subjects with vascular parkinsonism ( n = 10), essential tremor ( n = 7), progressive supranuclear palsy; PSP ( n = 7), multiple system atrophy; MSA ( n = 3), corticobasal syndrome; CBS ( n = 2), DLB ( n = 2), drug-induced tremor ( n = 2), dystonic tremor ( n = 2), restless legs syndrome ( n = 1), hemifacial spasm ( n = 1), motoneuron disease ( n = 1), amyotrophic shoulder neuralgia ( n = 1), and Alzheimer’s disease ( n = 1). The initial exploratory cohort consisted of ten PD subjects (8 men, mean age 67.1 ± 10.6) and ten healthy controls (5 men, mean age 65,7, SD ± 8,6.). For details, see Supplementary Table 3 ). The validation cohort included 99 PD subjects (49 men, mean age 66,1, SD ± 10,8), 36 healthy controls (20 men, mean age 63.7, SD ± 6,5.) and the described (see above) 41 OND subjects (29 men, mean age 70, SD ± 8.9. For details, see Supplementary Table 1 . The prodromal validation cohort consisted of 54 patients with iRBD (27 men, mean age 67.5, SD ± 8.1, for details, see Supplementary Table 4 ). RBD was diagnosed with two nights of state-of-the-art vPSG. Samples from HC were selected from the DeNoPa cohort 10 and matched for age and sex with the PD patients, had to be between 40 and 85 years old, without any active known/treated CNS condition, and with a negative family history of idiopathic PD. Antipsychotic drugs were an exclusion criterion. The provided data for sex are based on self-report.
The paired sample analysis of CSF, plasma and serum was applied in samples from subjects with OND 7 men, mean age 74 years, SD ± 7; diagnosis: four Alzheimer’s disease, three vascular Parkinsonism, one essential tremor, one multiple system atrophy one progressive supranuclear palsy).
Clinical assessments included the UPDRS subscores (parts I–III), the sum (UPDRS total score), and cognitive screening using the MMSE 10 .
Plasma and serum samples for both cohorts were collected in the morning under fasting conditions using Monovette tubes (Sarstedt, Nümbrecht, Germany) for EDTA plasma and serum collection by venipuncture. Tubes were centrifuged at 2500× g at room temperature (20 °C) for 10 min and aliquoted and frozen within 30 min of collection at −80 °C until analysis 10 , 66 . Single- use aliquots were used for all analyses presented here. For further details, we refer to the following publication 67 .
CSF was collected in polypropylene tubes (Sarstedt, Nümbrecht, Germany) directly after the plasma collection by lumbar puncture in the sitting position. Tubes were centrifuged at 2500× g at room temperature (20 °C) for 10 min and aliquoted and frozen within 30 min after collection at −80 °C until analysis. Before centrifugation, white and red blood cell counts in CSF were determined manually 10 , 66 . CSF β-amyloid 1–42, total tau protein (t-tau), phosphorylated tau protein (p-tau181) and neurofilament light chains (NFL) concentrations were measured by board-certified laboratory technicians, who were blinded to clinical data, using commercially available INNOTEST ELISA kits for the tau and Aβ markers (Fujirebio Europe, Ghent, Belgium) and the UmanDiagnostics NF-light® assay (UmanDiagnostics, Umeå, Sweden) for NFL. Total protein and albumin levels were measured by nephelometry (Dade Behring/Siemens Healthcare Diagnostics) 66 .
For the α-synuclein seeding aggregation assay (αSyn-SAA) the CSF samples were blindly analyzed in triplicate (40 μL/well) in a reaction mixture (0.3 mg/mL recombinant α-Syn (Amprion [California, USA]; catalogue number S2020), 100 mM piperazine- N , N ′-bis(2-ethanesulfonic acid) (PIPES) pH 6.50, 500 mM sodium chloride, 10 μM thioflavin T, and one bovine serum albumin (BSA)–blocked 2.4-mm silicon nitride G3 bead (Tsubaki-Nakashima [Georgia, USA]). Beads were blocked in 1% BSA 100 mM PIPES pH 6.50 and washed with 100 mM PIPES pH 6.50. The assay was performed in 96-well plates (Costar [New York, USA], catalogue number 3916) using a FLUOstar Omega fluorometer (BMG [Ortenberg, Germany]). Plates were orbitally shaken (800 rpm for 1 min every 29 min at 37 °C). Results from the triplicates were considered input for a three-output probabilistic algorithm with sample labelling as “positive,” “negative,” or “inconclusive”, based on the parameters: Maximum fluorescence (Fmax), time to reach 50% Fmax (T50), slope, and the coefficient of determination for the fitting were calculated for each replicate using a sigmoidal equation available in Mars data analysis software (BMG). The time to reach the 5000 relative fluorescence units (RFU) threshold (TTT) was calculated with a user-defined equation in Mars 19 .
Discovery plasma proteomics (phase 0)
In the mass spectrometry-based proteomic discovery analysis of plasma, we depleted the control and de novo PD samples from the twelve most abundant plasma proteins using Pierce Top12 columns (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. The depleted samples were freeze-dried before the addition of 20 µL of lysis buffer (100 mM Tris pH 7.8, 6 M urea, 2 M thiourea, and 2% ASB-14). The samples were shaken on an orbital shaker for 60 min at 1500 rpm. To break disulphide bonds, 45 µg DTE was added, and the samples were incubated for 60 min. To prevent disulphide bonds from reforming, 108 µg IAA was added, and the samples were incubated for 45 min covered in light. About 165 µL MilliQ water was added to dilute the concentration of urea and 1 µg trypsin gold (Promega, Mannheim, Germany) was added before 16 h of incubation at +37 °C to digest the proteins into peptides. To purify the peptides, solid phase extraction was performed using 100 mg C18 cartridges (Biotage, Uppsala, Sweden). The cartridges were washed with two 1 mL aliquots of 60% ACN, and 0.1% TFA before equilibration by two 1 mL aliquots of 0.1% TFA. The concentration of TFA in the samples was adjusted to 0.1%. The samples were loaded, and the flow-through was captured and re-applied. Salts were washed away from the bound peptides by two 1 mL aliquots of 0.1% TFA. The peptides were eluted by two 250 µL aliquots of 60% ACN, and 0.1% TFA. Solvents were evaporated using a vacuum concentrator. The samples were re-suspended in 50 µL 3% ACN, 0.1% FA prior to analysis. About 4 µL was injected into a 2D-NanoAquity liquid chromatography system (Waters, Manchester, UK). All samples were fractionated online into ten fractions over 12 h. The mobile phase in the first chromatographic system consisted of A1: 10 mM ammonium hydroxide titrated to pH 9 and B1: acetonitrile. The second chromatographic system’s mobile phase was A2: 5% dimethylsulfoxide (DMSO) + 0.1% formic acid, B2: acetonitrile with 5% DMSO + 0.1% formic acid. 2D-liquid chromatography fractionation was performed by loading the sample onto a 300 µm × 50 mm, 5 µm Peptide BEH C18 column (Waters). The peptides were eluted from the first column at a flow rate of 2 µL/min. The initial condition of the gradient elution was 3% B, held over 0.5 minutes before linearly increasing the proportion of organic solvent B, fraction per fraction over 0.5 min. The conditions thereafter remained static for 4 min before returning to the initial conditions over 0.5 min and equilibration prior to the next elution for 10 min. The eluted peptides from the first-dimensional column were loaded into a 180 µm × 20 mm, 5 µm Symmetry C18 trap column (Waters) before entering the analytical column, a 75 µm × 150 mm, 1.7 µm Peptide BEH C18 (Waters). The column temperature was +45 °C. The gradient elution applied to the analytical column started at 3% B and was linearly increased to 40% B over 40 min after which it was increased to 85% B over 2 min and washed for 2 min before returning to initial conditions over 2 min followed by equilibration for 15 min before the subsequent injection. The eluted peptides were detected using a Synapt-G2-S i (Waters) equipped with a nano-electrospray ion source. Data were acquired in positive MS E mode from 0 to 60 min within the m/z range 50−2000. The capillary voltage was set to 3 kV and the source temperature to +100 °C. The desolvation gas consisted of nitrogen with a flow rate of 50 L/h, and the desolvation temperature was set to +200 °C. The purge and desolvation gas consisted of nitrogen, operated at a flow rate of 600 mL/h and 600 L/h, respectively. The gas in the IMS cell was helium, with a flow rate of 90 mL/h. The low energy acquisition was performed by applying a constant collision energy of 4 V with a 1-s scan time. High energy acquisition was performed by applying a collision energy ramp, from 15 to 40 V, and the scan time was 1 s. The lock mass consisted of 500 fmol/µL [glu1]-fibrinopeptide B, continuously infused at a flow rate of 0.3 µL/min and acquired every 30 s. The doubly charged precursor ion, m/z 785.8426, was utilised for mass correction. After acquisition, data were imported to Progenesis QI for proteomics (Waters), and the individual fractions were processed before all results were merged into one experiment. The Ion Accounting workflow was utilised, with UniProt Canonical Human Proteome as a database (build 2016). The digestion enzyme was set as trypsin. Carbamidomethyl on cysteines was set as a fixed modification; deamidation of glutamine and asparagine, and oxidation of tryptophan and pyrrolidone carboxylic acid on the N-terminus were set as variable modifications. The identification tolerance was restricted to at least two fragments per peptide, three fragments per protein, and one peptide per protein. A FDR of 4% or less was accepted. The resulting identifications and intensities were exported and variables with a confidence score less than 15 and only one unique peptide were filtered out.
Targeted plasma proteomics (phase I)
The peptides included in the targeted assay were selected from several proteomic screening studies in which we analysed plasma, serum, urine, and CSF in ageing, PD and AD. The analytical method is described by ref. 17 . Furthermore, due to the suggested involvement of inflammation in neurodegenerative diseases, several known pro- and anti-inflammatory proteins identified from the literature were included in the multiplexed assay. The final panel consisted of 121 proteins (Supplementary Table 2 ), out of which a number were measured with two peptides, leading to a total of 167 unique peptides. When possible, the peptides were chosen to have an amino acid sequence length between 7 and 20. The amino acid sequences were confirmed to be unique to the proteins by using the Basic Local Alignment Search Tool (BLAST) provided by UniProt 68 . Synthetic peptide standards were purchased from GenScript (Amsterdam, Netherlands). To establish the most optimal transitions, repeated injections of 1 pmol peptide standard onto a Waters Acquity ultra-performance liquid chromatography (UPLC) system coupled to a Waters Xevo-TQ-S triple quadrupole MS were performed. The most high-abundant precursor-to-product ion transitions and their optimal collision energies were determined manually or using Skyline 69 . Detection was performed in positive ESI mode. The capillary voltage was set to 2.8 kV, the source temperature to 150 °C, the desolvation temperature to 600 °C, and the cone gas and desolvation gas flows to 150 and 1000 L/h, respectively. The collision gas consisted of nitrogen and was set to 0.15 mL/min. The nebuliser operated at 7 bar. Two transitions were chosen, one quantifier for relative concentration determination and one qualifier for identification, totally rendering 334 analyte transitions. Cone and collision energies varied depending on the optimal settings for each peptide. Each peptide was measured with a minimum of 12 points per peak and a dwell time of 10 ms or more to ensure adequate data acquisition. The optimised transitions were distributed over two multiple reaction monitoring (MRM) methods, always keeping the quantifier and qualifier for each peptide in the same MRM segment. Plasma, serum, and CSF samples were depleted from albumin and IgG using Pierce Top2 cartridges (Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer’s instructions. About 150 µg whole protein yeast enolase (ENO1) was added to the cartridges as an internal standard to account for digestion efficiency. Digestion was performed as described above. Solid phase extraction was carried out on BondElute 100 mg C18 96-well plates (Agilent, Santa Clara, USA) using the same methodology as in the preparation of untargeted proteomic analyses. Quality control samples were prepared from acetone-precipitated plasma, digested and solid phase extracted. Calibration curves ranging from 0 to 1 pmol/μL were constructed in blank and matrix by spiking increasing amounts of peptides into blank and QC samples. Before analysis, the samples were reconstituted in 30 µL 3% ACN, 0.1% FA containing 0.1 μM heavy isotope labelled peptides from the following proteins (annotated by gene name): ALDOA, C3, GSTO1, RSU1 and TSP1. About 5 µL were injected. The peptides were separated and detected on an Acquity UPLC system coupled to a Xevo-TQ-S triple quadrupole mass spectrometer (Waters, Manchester, UK). Chromatographic separation of the peptides was performed using a 1 × 100 mm, 1.7 μm ACQUITY UPLC Peptide CSH C18 column (Waters).
The mobile phase consisted of A: 0.1% formic acid and B: 0.1% formic acid in acetonitrile pumped at a flow rate of 0.2 mL/min. The column temperature was set to +55 °C. The initial mobile phase composition was 3% B, which was kept static for 0.8 min before initialising the linear gradient, running for 7.6 min to 25% B, eluting most of the peptides. B was thereafter linearly increased to 80% over 0.5 min and held for 1.9 min, eluting the most apolar peptides and washing the column before returning to the initial conditions over 0.1 minutes followed by equilibration for 6 min prior to the subsequent injection. Two subsequent injections of each sample were performed, each paired with one of the two MRM acquisition methods.
After acquisition, peak-picking and integration were performed using TargetLynx (version 4.1, Waters) or an in-house application ('mrmIntegrate') written in Python (version 3.8). mrmIntegrate is publicly available to download via the GitHub repository https://github.com/jchallqvist/mrmIntegrate . The application takes text files as input (.raw files are transformed into text files through the application 'MSConvert' from ProteoWizard 70 and applies a LOWESS filter over five points of the chromatogram. The integration method to produce areas under the curve is trapezoidal integration. The application enables retention time alignment and simultaneous integration of the same transition for all samples. Peptide peaks were identified by the blank and matrix calibration curves. The integrated peak areas were exported to Microsoft Excel, where first, the ratio between quantifier and qualifier peak areas were evaluated to ensure that the correct peaks had been integrated. The digestion efficiency was evaluated by monitoring the presence of baker’s yeast ENO1 in the samples, all samples without a signal were excluded from further analysis. After the initial quality assessment, the quantifier area was divided by the area of one of the internal standards, ALDOA or GSTO1 to yield a ratio used for the determination of relative concentrations. Any compound that also showed an intensity signal in the blank samples had the blank signal subtracted from the analyte peak intensity. Pooled plasma quality control samples were additionally evaluated to assess the robustness of the run.
Refined LC-MS/MS method (phase II)
The rapid and refined targeted proteomics LC-MS/MS method contained only peptides from the 31 proteins observed in the original targeted proteomics method (121 proteins). We utilised a Waters Acquity (UPLC) system coupled to a Waters Xevo-TQ-XS triple quadrupole operating in positive ESI mode. The column was an ACQUITY Premier Peptide BEH C18, 300 Å, 1.7 µm, maintained at 40 °C. The mobile phase was A: 0.1% formic acid in water, and B: 0.1% formic acid in acetonitrile. The gradient elution profile was initiated with 5% B and held for 0.25 min before linearly increasing to 40% B over 9.75 min to elute and separate the peptides. The column was washed for 1.6 min with 85% B before returning to the initial conditions and equilibrating for 0.4 min. The flow rate was 0.6 mL/min. The settings of the mass spectrometer and the peak-picking method were the same as described in the prior section. Baker’s yeast ENO1 was utilised to monitor digestion efficiency and as an internal standard.
Statistical methods
Most of the statistical analyses were performed in Python (version 3.8.5). The untargeted and targeted datasets were inspected for outliers and instrumental drift using principal component analysis (PCA) and orthogonal projection to latent variables (OPLS) in SIMCA, version 17 (Umetrics Sartorius Stedim, Umeå, Sweden). Outliers exceeding ten median deviations from each variable’s median were excluded. Instrumental drift was corrected by applying a non-parametric LOWESS filter from statsmodels (version 0.14.0) using 0.5 fractions of the data to estimate the LOWESS curve 71 . The data were evaluated for normal distribution using D’Agostino and Pearson’s method from SciPy (version 1.9.3) 72 . The non-normally distributed variables in the untargeted data were transformed to normality by the Box-Cox procedure using the SciPy function 'boxcox'. Significance testing between the independent groups of HC and PD/OND/iRBD individuals was performed by Student’s two-tailed t -test for the untargeted proteomic data and by Mann–Whitney’s non-parametric U -test (SciPy) for the targeted data. Due to the limited sample numbers, no multiple testing correction was performed in the untargeted data. In the targeted data, the Benjamini–Hochberg multiple testing correction procedure (statsmodels) was applied with an accepted false discovery rate of 5%. Fold-changes were calculated by dividing the means of the affected groups by the control group. Correlation analyses in the targeted data were performed by Spearman’s correlation (SciPy) and the correlation p values were adjusted variable-wise by the Benjamini–Hochberg procedure (FDR = 5%).
We implemented a support vector classifier model to discriminate between PD and HC and to predict new samples. The data were first z-scored protein-wise and any 'not a number'-values were replaced by the median. We used the 'LinearSVC' method from SciKit Learn and applied cross-validated recursive feature elimination to determine the number of variables to use in the model. The most discriminating variables for distinguishing between controls and PD were thereafter chosen by recursive feature elimination 73 . Feature selection and model training were performed on 70% of the data, partitioned using the SciKit Learn function “train_test_split”, and cross-validation was performed using a stratified k-fold with five splits. The remaining 30% of the data were predicted in the model. PR and ROC curves were constructed from the test data and consisted of each predictor and from the combined predictors, the packages precision_recall_curve and roc_curve from SciKit Learn were implemented. Linear mixed models were performed using the R-to-Python bridge software pymer4 (version 0.8.0), where individual was set as a random effect and the correlations between the MS measured proteins and clinical variables were evaluated for significance post Benjamini–Hochberg’s procedure for multiple testing correction. Plots of the data were constructed using the Seaborn and Matplotlib packages (versions 0.12.2 and 3.6.0, respectively) 74 .
All multivariate analyses were performed in SIMCA, version 17. OPLS and OPLS-discriminant analysis (OPLS-DA) models were evaluated for significance by ANOVA p values and by permutation tests applying 1000 permutations, where p < 0.05 and p < 0.001 were deemed significant, respectively.
Data were analysed for pathway enrichment using IPA (QIAGEN Inc. Data were analysed for pathway enrichment using IPA (QIAGEN Inc., https://digitalinsights.qiagen.com/products-overview/discovery-insights-portfolio/analysis-and-visualization/qiagen-ipa/ .). Input variables were set to proteins demonstrating a significant difference between PD individuals and HC, with fold-change as expression observation. The accepted output pathways were restricted to p < 0.05 and at least two proteins were included in the pathways. Gene Ontology (GO) annotations were extracted using DAVID Bioinformatics Resources (2021 build) 75 , 76 . Networks were built in Cytoscape 77 (version 3.8.0) by applying the “Organic layout” from yFiles 77 .
Obtaining biological materials
Patient samples can be provided to other researchers for certain projects after contact with the corresponding authors and upon availability approval of the team in Kassel, Germany.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The chromatograms from the targeted mass spectrometric data generated in this study have been deposited in the ProteomeXchange database under accession code PXD041419 and in the Panorama repository ( https://panoramaweb.org/DNP_Pub.url , https://doi.org/10.6069/p9cy-h335 ). The integrated targeted mass spectrometric data generated in this study are provided in the Supplementary Information. Source data for all data presented in graphs within the figures are provided in a source data file. Source data are provided with this paper.
Code availability
Peak-picking and integrations were performed in TargetLynx (part of the MassLynx suite, version 4.1), or using an in-house application written in Python which can be found on GitHub ( https://github.com/jchallqvist/mrmIntegrate ). The data visualisation and statistical analyses were performed in Python (version 3.8.5) using the packages SciPy (version 1.9.3), statsmodels (version 0.14.0), SciKit Learn (version 1.1.2), Seaborn (version 13.0) and Matplotlib (version 3.6.0). The code used can be found on GitHub ( https://github.com/jchallqvist/DNP_Pub/blob/main/DNP_Code , https://doi.org/10.5281/zenodo.11130369 ).
Simuni, T. et al. Baseline prevalence and longitudinal evolution of non-motor symptoms in early Parkinson’s disease: the PPMI cohort. J. Neurol. Neurosurg. Psychiatry 89 , 78–88 (2018).
Article PubMed Google Scholar
Michell, A. W., Lewis, S. J., Foltynie, T. & Barker, R. A. Biomarkers and Parkinson’s disease. Brain 127 , 1693–1705 (2004).
Article CAS PubMed Google Scholar
Kieburtz, K., Katz, R., McGarry, A. & Olanow, C. W. A new approach to the development of disease-modifying therapies for PD; fighting another pandemic. Mov. Disord. 36 , 59–63 (2021).
Shahnawaz, M. et al. Development of a biochemical diagnosis of Parkinson disease by detection of α-synuclein misfolded aggregates in cerebrospinal fluid. JAMA Neurol. 74 , 163–172, (2017).
Mollenhauer, B. et al. Validation of serum neurofilament light chain as a biomarker of Parkinson’s disease progression. Mov. Disord . https://doi.org/10.1002/mds.28206 (2020).
Lindestam Arlehamn, C. S. et al. α-Synuclein-specific T cell reactivity is associated with preclinical and early Parkinson’s disease. Nat. Commun. 11 , 1875 (2020).
Article ADS CAS PubMed PubMed Central Google Scholar
Mollenhauer, B. et al. Baseline predictors for progression 4 years after Parkinson’s disease diagnosis in the De Novo Parkinson Cohort (DeNoPa). Mov. Disord. 34 , 67–77 (2019).
Bartl, M. et al. Blood markers of inflammation, neurodegeneration, and cardiovascular risk in early Parkinson’s disease. Mov. Disord . https://doi.org/10.1002/mds.29257 (2022).
Simuni, T. et al. A biological definition of neuronal α-synuclein disease: towards an integrated staging system for research. Lancet Neurol. 23 , 178–190 (2024).
Mollenhauer, B. et al. Nonmotor and diagnostic findings in subjects with de novo Parkinson disease of the DeNoPa cohort. Neurology 81 , 1226–1234 (2013).
Hällqvist, J. et al. A multiplexed urinary biomarker panel has potential for Alzheimer’s disease diagnosis using targeted proteomics and machine learning. Int. J. Mol. Sci . https://doi.org/10.3390/ijms241813758 (2023).
Hu, W., Ralay Ranaivo, H., Craft, J. M., Van Eldik, L. J. & Watterson, D. M. Validation of the neuroinflammation cycle as a drug discovery target using integrative chemical biology and lead compound development with an Alzheimer’s disease-related mouse model. Curr. Alzheimer Res. 2 , 197–205 (2005).
Notter, T. et al. Translational evaluation of translocator protein as a marker of neuroinflammation in schizophrenia. Mol. Psychiatry 23 , 323–334 (2018).
Jonsson, M., Gerdle, B., Ghafouri, B. & Backryd, E. The inflammatory profile of cerebrospinal fluid, plasma, and saliva from patients with severe neuropathic pain and healthy controls-a pilot study. BMC Neurosci. 22 , 6 (2021).
Article PubMed PubMed Central Google Scholar
Chen, X. et al. Progranulin does not bind tumor necrosis factor (TNF) receptors and is not a direct regulator of TNF-dependent signaling or bioactivity in immune or neuronal cells. J. Neurosci. 33 , 9202–9213 (2013).
Article CAS PubMed PubMed Central Google Scholar
Captur, G. et al. Plasma proteomic signature predicts who will get persistent symptoms following SARS-CoV-2 infection. EBioMedicine 85 , 104293 (2022).
Doykov, I. et al. The long tail of Covid-19’ - The detection of a prolonged inflammatory response after a SARS-CoV-2 infection in asymptomatic and mildly affected patients. F1000Res 9 , 1349 (2020).
Hu, M. T. REM sleep behavior disorder (RBD). Neurobiol. Dis. 143 , 104996 (2020).
Concha-Marambio, L. et al. Accurate detection of α-synuclein seeds in cerebrospinal fluid from isolated rapid eye movement sleep behavior disorder and patients with Parkinson’s disease in the de novo Parkinson (DeNoPa) cohort. Mov. Disord. 38 , 567–578 (2023).
Dayon, L. et al. Proteomes of paired human cerebrospinal fluid and plasma: relation to blood-brain barrier permeability in older adults. J. Proteome Res. 18 , 1162–1174 (2019).
Whelan, C. D. et al. Multiplex proteomics identifies novel CSF and plasma biomarkers of early Alzheimer’s disease. Acta Neuropathol. Commun. 7 , 169 (2019).
Bloem, B. R., Okun, M. S. & Klein, C. Parkinson’s disease. Lancet 397 , 2284–2303 (2021).
Dorsey, E. R., Sherer, T., Okun, M. S. & Bloem, B. R. The emerging evidence of the Parkinson pandemic. J. Parkinsons Dis. 8 , S3–s8 (2018).
Grossauer, A. et al. α-Synuclein seed amplification assays in the diagnosis of synucleinopathies using cerebrospinal fluid-A systematic review and meta-analysis. Mov. Disord. Clin. Pr. 10 , 737–747 (2023).
Article Google Scholar
Okuzumi, A. et al. Propagative α-synuclein seeds as serum biomarkers for synucleinopathies. Nat. Med. 29 , 1448–1455 (2023).
Raffield, L. M. et al. Comparison of proteomic assessment methods in multiple cohort studies. Proteomics 20 , e1900278 (2020).
Hernández, B., Parnell, A. & Pennington, S. R. Why have so few proteomic biomarkers “survived” validation? (Sample size and independent validation considerations). Proteomics 14 , 1587–1592 (2014).
Füzéry, A. K., Levin, J., Chan, M. M. & Chan, D. W. Translation of proteomic biomarkers into FDA approved cancer diagnostics: issues and challenges. Clin. Proteom. 10 , 13 (2013).
Bader, J. M., Albrecht, V. & Mann, M. MS-based proteomics of body fluids: the end of the beginning. Mol. Cell Proteom. 22 , 100577 (2023).
Article CAS Google Scholar
Pan, C. et al. Targeted discovery and validation of plasma biomarkers of Parkinson’s disease. J. Proteome Res. 13 , 4535–4545 (2014).
Qin, X. Y., Zhang, S. P., Cao, C., Loh, Y. P. & Cheng, Y. Aberrations in peripheral inflammatory cytokine levels in Parkinson disease: a systematic review and meta-analysis. JAMA Neurol. 73 , 1316–1324, (2016).
Choi, M. L. & Gandhi, S. Crucial role of protein oligomerization in the pathogenesis of Alzheimer’s and Parkinson’s diseases. FEBS J. 285 , 3631–3644 (2018).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334 , 1081–1086 (2011).
Article ADS CAS PubMed Google Scholar
Bertolotti, A., Zhang, Y. H., Hendershot, L. M., Harding, H. P. & Ron, D. Dynamic interaction of BiP and ER stress transducers in the unfolded-protein response. Nat. Cell Biol. 2 , 326–332 (2000).
Colla, E. Linking the endoplasmic reticulum to Parkinson’s disease and alpha-synucleinopathy. Front. Neurosci. 13 , 560 (2019).
Mercado, G., Castillo, V., Soto, P. & Sidhu, A. ER stress and Parkinson’s disease: pathological inputs that converge into the secretory pathway. Brain Res. 1648 , 626–632 (2016).
Loeffler, D. A., Camp, D. M. & Conant, S. B. Complement activation in the Parkinson’s disease substantia nigra: an immunocytochemical study. J. Neuroinflamm 3 , 29 (2006).
Zattoni, M. et al. Serpin signatures in Prion and Alzheimer’s diseases. Mol. Neurobiol. 59 , 3778–3799 (2022).
Seo, M. H. & Yeo, S. Association of increase in Serping1 level with dopaminergic cell reduction in an MPTP-induced Parkinson’s disease mouse model. Brain Res. Bull. 162 , 67–72 (2020).
Anderson, J. P. et al. Phosphorylation of Ser-129 is the dominant pathological modification of alpha-synuclein in familial and sporadic Lewy body disease. J. Biol. Chem. 281 , 29739–29752 (2006).
Fujiwara, H. et al. alpha-Synuclein is phosphorylated in synucleinopathy lesions. Nat. Cell Biol. 4 , 160–164 (2002).
Kim, K. S. et al. Proteolytic cleavage of extracellular alpha-synuclein by plasmin implications for Parkinson disease. J. Biol. Chem. 287 , 24862–24872 (2012).
Reho, P. et al. GRN mutations are associated with Lewy body dementia. Mov. Disord. 37 , 1943–1948 (2022).
Kao, A. W., Mckay, A., Singh, P. P., Brunet, A. & Huang, E. J. Progranulin, lysosomal regulation and neurodegenerative disease. Nat. Rev. Neurosci. 18 , 325–333 (2017).
Mateo, I. et al. Reduced serum progranulin level might be associated with Parkinson’s disease risk. Eur. J. Neurol. 20 , 1571–1573 (2013).
Bachmann-Harildstad, G. Diagnostic values of beta-2 transferrin and beta-trace protein as markers for cerebrospinal fluid fistula. Rhinology 46 , 82–85 (2008).
PubMed Google Scholar
Pediaditakis, I. et al. Modeling alpha-synuclein pathology in a human brain-chip to assess blood-brain barrier disruption. Nat. Commun. 12 , 5907 (2021).
Serafino, A., Giovannini, D., Rossi, S. & Cozzolino, M. Targeting the Wnt/β-catenin pathway in neurodegenerative diseases: recent approaches and current challenges. Expert Opin. Drug Discov. 15 , 803–822 (2020).
Arenas, E. Wnt signaling in midbrain dopaminergic neuron development and regenerative medicine for Parkinson’s disease. J. Mol. Cell Biol. 6 , 42–53 (2014).
L’Episcopo, F. et al. A Wnt1 regulated Frizzled-1/β-Catenin signaling pathway as a candidate regulatory circuit controlling mesencephalic dopaminergic neuron-astrocyte crosstalk: therapeutical relevance for neuron survival and neuroprotection. Mol. Neurodegener. 6 , 49 (2011).
Sweeney, M. D., Sagare, A. P. & Zlokovic, B. V. Blood-brain barrier breakdown in Alzheimer disease and other neurodegenerative disorders. Nat. Rev. Neurol. 14 , 133–150 (2018).
L’Episcopo, F. et al. Wnt/beta-catenin signaling is required to rescue midbrain dopaminergic progenitors and promote neurorepair in ageing mouse model of Parkinson’s disease. Stem Cells 32 , 2147–2163 (2014).
Marchetti, B. Wnt/beta-catenin signaling pathway governs a full program for dopaminergic neuron survival, neurorescue and regeneration in the MPTP mouse model of Parkinson’s disease. Int. J. Mol. Sci. 19 , 3743 (2018).
Marchetti, B. et al. Parkinson’s disease, aging and adult neurogenesis: Wnt/beta-catenin signalling as the key to unlock the mystery of endogenous brain repair. Aging Cell 19 , e1310110 (2020).
L’Episcopo, F. et al. Neural stem cell grafts promote astroglia-driven neurorestoration in the aged Parkinsonian brain via Wnt/beta-catenin signaling. Stem Cells 36 , 1179–1197 (2018).
Serafino, A. et al. Developing drugs that target the Wnt pathway: recent approaches in cancer and neurodegenerative diseases. Expert Opin. Drug Discov. 12 , 169–186 (2017).
Harms, A. S., Ferreira, S. A. & Romero-Ramos, M. Periphery and brain, innate and adaptive immunity in Parkinson’s disease. Acta Neuropathol. 141 , 527–545 (2021).
Dong, M. X. et al. Serum butyrylcholinesterase activity: a biomarker for Parkinson’s disease and related dementia. Biomed. Res. Int. 2017 , 1524107 (2017).
Postuma, R. B. et al. Risk and predictors of dementia and parkinsonism in idiopathic REM sleep behaviour disorder: a multicentre study. Brain 142 , 744–759 (2019).
Zhang, H. et al. Risk factors for phenoconversion in rapid eye movement sleep behavior disorder. Ann. Neurol. 91 , 404–416 (2022).
Palma, J. A. et al. Prevalence of REM sleep behavior disorder in multiple system atrophy: a multicenter study and meta-analysis. Clin. Auton. Res. 25 , 69–75 (2015).
Jennings, D. et al. Imaging prodromal Parkinson disease: the Parkinson Associated Risk Syndrome Study. Neurology 83 , 1739–1746 (2014).
Schade, S. et al. Identifying prodromal NMS in a population-based recruitment strategy: Kassel data of Healthy Brain Ageing. Zenodo (2023).
Schade, S. et al. Acute levodopa challenge test in patients with de novo Parkinson’s disease: data from the DeNoPa cohort. Mov. Disord. Clin. Pr. 4 , 755–762 (2017).
Hummel, T., Sekinger, B., Wolf, S. R., Pauli, E. & Kobal, G. ‘Sniffin’ sticks’: olfactory performance assessed by the combined testing of odor identification, odor discrimination and olfactory threshold. Chem. Senses 22 , 39–52 (1997).
Mollenhauer, B. et al. Monitoring of 30 marker candidates in early Parkinson disease as progression markers. Neurology 87 , 168–177 (2016).
Mollenhauer, B. et al. α-Synuclein and tau concentrations in cerebrospinal fluid of patients presenting with parkinsonism: a cohort study. Lancet Neurol. 10 , 230–240 (2011).
UniProt. BLAST https://www.uniprot.org/blast/ . Accessed January 2024.
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26 , 966–968 (2010).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30 , 918–920 (2012).
Seabold, S. & Perktold, J. Statsmodels: econometric and statistical modeling with Python. In Proc. 9th Python in Science Conference (SciPy, 2010).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17 , 261–272 (2020).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12 , 2825–2830 (2011).
MathSciNet Google Scholar
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9 , 90–95 (2007).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50 , W216–221, (2022).
Huang Da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4 , 44–57 (2009).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13 , 2498–2504 (2003).
Download references
Acknowledgements
This work was supported by the Michael J Fox Foundation, PDUK, The Peto Foundation, The TMSRG (UCL), The BRC at Great Ormond Street Hospital, and the Horizon 2020 Framework Programme (Grant number 634821, PROPAG-AGING). We thank the PROPAG-AGING consortium, a full list of the members can be found in the supplementary material.
Open Access funding enabled and organized by Projekt DEAL.
Author information
These authors contributed equally: Jenny Hällqvist, Michael Bartl.
These authors jointly supervised this work: Kevin Mills, Brit Mollenhauer.
Authors and Affiliations
UCL Institute of Child Health and Great Ormond Street Hospital, London, UK
Jenny Hällqvist, Ivan Doykov, Justyna Śpiewak, Héloїse Vinette & Wendy E. Heywood
UCL Queen Square Institute of Neurology, Clinical and Movement Neurosciences, London, UK
Jenny Hällqvist & Kevin Mills
Department of Neurology, University Medical Center Goettingen, Goettingen, Germany
Michael Bartl, Mohammed Dakna, Mary Xylaki, Sandrina Weber & Brit Mollenhauer
Institute for Neuroimmunology and Multiple Sclerosis Research, University Medical Center Goettingen, Goettingen, Germany
Michael Bartl
Paracelsus-Elena-Klinik, Kassel, Germany
Sebastian Schade, Maria-Lucia Muntean, Friederike Sixel-Döring, Claudia Trenkwalder & Brit Mollenhauer
Department of Experimental, Diagnostic, and Specialty Medicine (DIMES), University of Bologna, Bologna, Italy
Paolo Garagnani, Chiara Pirazzini & Claudio Franceschi
IRCCS Istituto delle Scienze Neurologiche di Bologna, Bologna, Italy
Maria-Giulia Bacalini
National Hospital for Neurology & Neurosurgery, Queen Square, WC1N3BG, London, UK
Kailash Bhatia & Sebastian Schreglmann
Institute of Diagnostic and Interventional Neuroradiology, University Medical Center Goettingen, Goettingen, Germany
Marielle Ernst
Department of Neurology, Philipps-University, Marburg, Germany
Friederike Sixel-Döring
UCL: Food, Microbiomes and Health Institute Strategic Programme, Quadram Institute Bioscience, Norwich Research Park, Norwich, UK
Héloїse Vinette
Department of Neurosurgery, University Medical Center Goettingen, Goettingen, Germany
Claudia Trenkwalder
You can also search for this author in PubMed Google Scholar
Contributions
J.H., M.B., K.M., and B.M. conceptualised, planned and oversaw all aspects of the study. J.H., K.M., J.S., H.V., M.B. and S. Schreglmann performed and analyzed most of the experiments. S. Schade, S.W. and M.B. consented to the subjects and collected the samples. M.-L.M., F.S.-D. and S. Schade analyzed the sleep lab data and diagnosed the iRBD subjects. J.H. and M.D. performed the statistical data analysis. J.H. applied the machine learning methods and designed the figures. W.H., I.D., C.F., M.-G.B., P.G., C.P., K.B. and M.X. provided substantial contributions to the conception of the work, acquisition and interpretation of the data, particularly for the mass spectrometry setup and the refinement of the targeted panel. S. Schade, S.W., C.T., M.B., B.M., M.-L.M. and F.S.D. conceptualised the clinical study, analyzed the clinical data and reevaluated the diagnosis. M.E. provided substantial contributions to the clinical data analyzes, particularly the imaging patient data in regard to differential diagnosis. J.H., M.B., K.M. and B.M. wrote the manuscript with input and substantial revisions from all authors.
Corresponding authors
Correspondence to Jenny Hällqvist or Michael Bartl .
Ethics declarations
Competing interests.
JH, MD, MX, SW, KB, ME, PG, MGB, CP, KM, ID, WH, JS, HV and CF and have no competing interests to report. MB has received funding from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – 413,501,650. CT has received honoraria for consultancy from Roche, and honoraria for educational lectures from UCB, and has received research funding for the PPMI study from the Michael J. Fox Foundation and funding from the EU (Horizon 2020) and stipends from the (International Parkinson’s and Movement Disorder Society) IPMDS. BM has received honoraria for consultancy from Roche, Biogen, AbbVie, UCB, and Sun Pharma Advanced Research Company. BM is a member of the executive steering committee of the Parkinson Progression Marker Initiative and PI of the Systemic Synuclein Sampling Study of the Michael J. Fox Foundation for Parkinson’s Research and has received research funding from the Deutsche Forschungsgemeinschaft (DFG), EU (Horizon 2020), Parkinson Fonds Deutschland, Deutsche Parkinson Vereinigung, Parkinson’s Foundation and the Michael J. Fox Foundation for Parkinson’s Research. MLM has received honoraria for speaking engagements from Deutsche Parkinson Gesellschaft e.V., and royalties from Gesellschaft fur Medien + Kommunikation mbH + Co. FSD has received honoraria for speaking engagements from AbbVie, Bial, Ever Pharma, Medtronic and royalties from Elsevier and Springer. She served on an advisory board for Zambon and Stada Pharma. FSD participated in Ad Boards for consultation: Abbvie, UCB, Bial, Ono, Roche and got honorary for lecturing: Stada Pharm, AbbVie, Alexion, Bial. S. Schade received institutional salaries supported by the EU Horizon 2020 research and innovation programme under grant agreement No. 863664 and by the Michael J. Fox Foundation for Parkinson’s Research under grant agreement No. MJFF-021923. He is supported by a PPMI Early Stage Investigators Funding Programme fellowship of the Michael J. Fox Foundation for Parkinson’s Research under grant agreement No. MJFF-022656. S. Schreglmann received institutional salaries supported by the EU Horizon 2020 research and innovation programme under grant agreement No. 863664, support from the Advanced Clinician Scientist programme by the Interdisciplinary Centre for Clinical Research, Wuerzburg, Germany, and from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Project-ID 424778381-TRR 295. He is a fellow of the Thiemann Foundation. He serves as a scientific adviser to Elemind Inc.
Peer review
Peer review information.
Nature Communications thanks Kanta Horie and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information, peer review file, reporting summary, source data, source data, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Hällqvist, J., Bartl, M., Dakna, M. et al. Plasma proteomics identify biomarkers predicting Parkinson’s disease up to 7 years before symptom onset. Nat Commun 15 , 4759 (2024). https://doi.org/10.1038/s41467-024-48961-3
Download citation
Received : 06 April 2023
Accepted : 20 May 2024
Published : 18 June 2024
DOI : https://doi.org/10.1038/s41467-024-48961-3
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing: Translational Research newsletter — top stories in biotechnology, drug discovery and pharma.
| Acta Crystallographica Section D Acta Crystallographica Section D STRUCTURAL BIOLOGY |
1. Introduction
2. materials and methods, 4. discussion, 5. conclusion and outlook, supporting information.

| Format | BIBTeX | |
| EndNote | ||
| RefMan | ||
| Refer | ||
| Medline | ||
| CIF | ||
| SGML | ||
| Plain Text | ||
| Text | ||

research papers \(\def\hfill{\hskip 5em}\def\hfil{\hskip 3em}\def\eqno#1{\hfil {#1}}\)
| STRUCTURAL BIOLOGY |

Factors affecting macromolecule orientations in thin films formed in cryo-EM
a National Centre for Biological Sciences, Tata Institute of Fundamental Research, GKVK Post, Bellary Road, Bengaluru 560 065, India * Correspondence e-mail: [email protected]
The formation of a vitrified thin film embedded with randomly oriented macromolecules is an essential prerequisite for cryogenic sample electron microscopy. Most commonly, this is achieved using the plunge-freeze method first described nearly 40 years ago. Although this is a robust method, the behaviour of different macromolecules shows great variation upon freezing and often needs to be optimized to obtain an isotropic, high-resolution reconstruction. For a macromolecule in such a film, the probability of encountering the air–water interface in the time between blotting and freezing and adopting preferred orientations is very high. 3D reconstruction using preferentially oriented particles often leads to anisotropic and uninterpretable maps. Currently, there are no general solutions to this prevalent issue, but several approaches largely focusing on sample preparation with the use of additives and novel grid modifications have been attempted. In this study, the effect of physical and chemical factors on the orientations of macromolecules was investigated through an analysis of selected well studied macromolecules, and important parameters that determine the behaviour of proteins on cryo-EM grids were revealed. These insights highlight the nature of the interactions that cause preferred orientations and can be utilized to systematically address orientation bias for any given macromolecule and to provide a framework to design small-molecule additives to enhance sample stability and behaviour.
Keywords: cryo-EM ; thin films ; preferred macromolecular orientation ; surfactants ; temperature .
EMDB references: CRP pentamer with CTAB, EMD-37864 ; CRP decamer with CTAB, EMD-37865 ; PaaZ with CTAB at 4°C, EMD-37866 ; catalase with SLS, EMD-37952 ; spike with CTAB, EMD-37953 ; catalase at 20°C, EMD-37954 ; catalase at 4°C, EMD-37955 ; catalase with CTAB, EMD-37956 ; β-galactosidase, no tag, EMD-39808 ; β-galactosidase, with tag, EMD-39809
PDB references: CRP pentamer with CTAB, 8wv4 ; CRP decamer with CTAB, 8wv5 ; PaaZ with CTAB at 4°C, 8wv6 ; catalase with SLS, 8wzh ; spike with CTAB, 8wzi ; catalase at 20°C, 8wzj ; catalase at 4°C, 8wzk ; catalase with CTAB, 8wzm
| Examples of anisotropic cryo-EM maps resulting from orientation bias. The upper panel shows the reference-free 2D class averages of ( ) SARS-CoV-2 spike protein and ( ) human erythrocyte catalase. For the spike protein, preferred bottom views are observed. In the case of catalase, a preference for the top/bottom view is evident. In the lower panel, 3D maps with anisotropic features are shown for the preferred and perpendicular views as labelled. The symmetries applied during reconstruction were 1 and 2 for the spike protein and catalase, respectively. |
To achieve this goal, we tested some commonly used surfactants with different properties on a set of five proteins: C-reactive protein (CRP) pentamers, CRP decamers, catalase, PaaZ and spike. In addition, we explored the effect of the presence of the histidine tag for spike and β -galactosidase and of physical factors such as the temperature during the sample-application step for catalase and PaaZ. We also serendipitously observed an effect of the grid hole dimensions of the holey carbon grid on the orientation distribution of catalase and discuss this briefly. Through this analysis, we identified factors that affect and determine the behaviour of the macromolecule on grids before freezing and studied their effects with a focus on the preferred orientation problem. This account highlights the factors that contribute to orientation bias and provides valuable information that can assist in achieving the optimal freezing conditions for any given macromolecule.
2.1. Source of proteins
Human C-reactive protein (catalogue No. C4063) and human erythrocyte catalase (catalogue No. C3556) were obtained from Sigma–Aldrich. The protein samples were either concentrated using an Amicon 100 kDa concentrator or diluted in respective buffers for grid freezing. All detergent stocks were made in ultrapure water and dilutions were made and used on the day of the experiment.
The SARS-CoV-2 S plasmid was a kind gift from the Krammer laboratory at Icahn School of Medicine, Mount Sinai. The spike gene was amplified from the plasmid and subcloned in the BacMam vector with a C-terminal HRV 3C cleavage tag followed by a seven-histidine and twin Strep tag. Bacmid DNA and virus were prepared as described in the Invitrogen Bac-to-Bac manual. After two generations of amplification in Sf9 cells, the V2 virus was used for transfection of HEK293F cells at a density of 2 million per millilitre. Sodium butyrate (4 m M ) was added to enhance the production of protein 8 h post-infection. The medium supernatant containing the secreted spike protein was harvested on day 3 by centrifuging the cells at 150 g for 10 min. The medium was incubated with pre-equilibrated Ni–NTA (Qiagen) beads at room temperature for 1–2 h (1 ml of beads per 200 ml of medium). The Ni–NTA beads were washed with phosphate-buffered saline (PBS) containing 20 m M imidazole, followed by elution with 280 m M imidazole in PBS. The eluted protein was run on SDS–PAGE to assess its purity, further concentrated and injected onto a 24 ml Superdex 200 (Cytiva) size-exclusion column to exchange the buffer to 50 m M Tris pH 8, 200 m M NaCl, 1 m M DTT. To cleave the tag, the eluted fractions from Ni–NTA chromatography were diluted with 50 m M Tris pH 8, 200 m M NaCl, 1 m M DTT and incubated with HRV 3C protease overnight at 4°C, followed by reverse IMAC to obtain the spike protein without tag in the flowthrough. The flowthrough was concentrated using an Amicon 100 kDa concentrator, flash-frozen using liquid nitrogen and stored at −80°C until further use.
2.2. Grid preparation
6.3 µl of the protein was thawed on ice and 0.7 µl of 10× additive (surfactant) stock was added to obtain a final concentration of 1×. This sample was incubated on ice for 2–5 min and then centrifuged at 21 000 g for 20 min. Meanwhile, a Vitrobot Mark IV (Thermo Fisher Scientific) chamber was equilibrated at 20°C (unless stated otherwise) and 100% humidity. Quantifoil 1.2/1.3 or Quantifoil 0.6/1 grids were glow-discharged in a reduced-air environment with a PELCO easiGlow chamber using a standard setting of 25 mA current for 1 min. The grid was mounted on the Vitrobot Mark IV and 3 µl of sample was applied to the grid. A blotting time of 3–4 s, a wait time of 10 s and a blot force of 0 were used to obtain a thin film of the specimen. For data sets where grids were prepared at different temperatures, the protein was incubated on a thermal block at the required temperature for 3–7 min before applying it to the grid. The Vitrobot chamber was maintained at the required temperature and 100% humidity. The blot time, blot force and wait time were kept constant.
IMAGES
VIDEO
COMMENTS
The acknowledgement section of a research paper should include acknowledgements for individuals or entities that have contributed to the research process. This may include mentors, advisors, funding agencies, research collaborators, or others who have provided valuable support, guidance, or resources.
Formatting the acknowledgements. As a rule of thumb, the acknowledgement section should be a single short paragraph of say half a dozen lines. Examine the target journal for the format: whether the heading appears on a separate line or run on (that is, the text follows the heading on the same line). Check also whether the heading is in bold or ...
The acknowledgements section is your opportunity to thank those who have helped and supported you personally and professionally during your thesis or dissertation process. Thesis or dissertation acknowledgements appear between your title page and abstract and should be no longer than one page. In your acknowledgements, it's okay to use a more ...
Writing the acknowledgements section of your thesis or dissertation is an opportunity to express gratitude to everyone who helped you along the way. Remember to: Acknowledge those people who significantly contributed to your research journey. Order your thanks from formal support to personal support. Maintain a balance between formal and ...
Acknowledgement in Research Paper - A Quick Guide [5 Examples] Academic. The acknowledgement section in your research paper is where you thank those who have helped or supported you throughout your research and writing. It is a short section of 3-5 paragraphs or no more than 300 words you put on a page after the title page.
Acknowledgements 101: The Basics. The acknowledgements section in your thesis or dissertation is where you express gratitude to those who helped bring your project to fruition. This section is typically brief (a page or less) and less formal, but it's crucial to thank the right individuals in the right order.. As a rule of thumb, you'll usually begin with academic support: your supervisors ...
Writing acknowledgements in a research paper is a thoughtful process that requires careful consideration of who to acknowledge and how to express gratitude. By following the guidelines and examples provided in this article, you can write effective acknowledgements that reflect your appreciation and respect for the contributions of others to ...
Acknowledgments. This usually follows the Discussion and Conclusions sections. Its purpose is to thank all of the people who helped with the research but did not qualify for authorship (check the target journal's Instructions for Authors for authorship guidelines). Acknowledge anyone who provided intellectual assistance, technical help ...
The Purpose of Acknowledgements. The acknowledgement section of a thesis or dissertation is where you recognise and thank those who supported you during your PhD. This can be but is not limited to individuals, institutions or organisations. Although your acknowledgements will not be used to evaluate your work, it is still an important section of your thesis.
My labmate's thesis, who wrote the acknowledgements in a different style to the rest by using bullet points. Dr Wei's thesis, acknowledgements on page 6. Direct download here. Shortest acknowledgements section of the list at 122 words. Dr Manca's thesis, acknowledgements on page 5. Direct download here. Dr Liu's thesis, page 5.
There are several ways you could word your thanks. A few suggestions are: This research would not have been possible without…. My sincere thanks go to…. I am grateful to…. Heartfelt thanks to…. I would like to express my gratitude to…. Appreciation is due to…. I acknowledge the contribution of….
📋 Where do you put acknowledgements in a research paper? While acknowledgments are usually more present in academic theses, they can also be a part of research papers. In academic theses, acknowledgments are usually found at the beginning, somewhere between abstract and introduction. In research papers, acknowledgments are usually found at ...
help you write your Acknowledgements section of your dissertation. According to one source, the Acknowledgements section of a Ph.D. dissertation is the most widely read section. Whether you believe this or not, many individuals who helped you in the process of writing may check to see if, indeed, they have meant something to you.
Tips on Writing Acknowledgement for Thesis. Sincerity is Key: Ensure your acknowledgment reflects genuine gratitude. Be Specific: Mention names and their roles clearly. Professional Tone: Maintain a formal but heartfelt tone. Brevity Matters: Keep it concise, focusing on key contributors.
The acknowledgment section helps identify the contributors responsible for specific parts of the project. It can include: Authors. Non-authors (colleagues, friends, supervisor, etc.) Funding sources. Editing services, Administrative staff. In academic writing, the information presented in the acknowledgment section should be kept brief.
The acknowledgement section in a dissertation is used to express gratitude towards all those who have helped you prepare the dissertation. Both professional and personal acknowledgements can be included in it. The acknowledgement section comes in between the title page and the abstract page. It is best suited to be on one page.
Finally, the ICMJE encourages written permission from acknowledged individuals "because acknowledgment may imply endorsement.". Funding sources should also be mentioned in the acknowledgments section, unless your target journal requires a separate section for this information. Whether the funding was partial or full, relevant grant numbers ...
The Acknowledgments section is present in both a paper and an academic thesis. For papers, the Acknowledgments section is usually presented at the back, whereas in a thesis, this section is located towards the front of the manuscript and is commonly placed somewhere between the abstract and Introduction.
Acknowledgement in a thesis serves as a gesture of respect and gratitude towards those who have contributed to the student's success. It is a way of recognizing the support and guidance received throughout the research and writing process. By acknowledging the contributions of others, the student acknowledges the collective effort that has ...
Acknowledgement for paper publication is a crucial aspect that recognizes the contributions and support received throughout the research process. It holds significant value in academic papers, showcasing appreciation for the individuals and institutions that have played a role in the study's success.
It is common to include an acknowledgement section in the beginning of a research paper, usually after the abstract and before the main body of the paper. The acknowledgement should be brief and to the point, and it is generally written in a formal, professional tone. You should avoid using casual language or personal anecdotes in this section.
Get white paper. Take preventative measures against network degradation. "With service-centric assurance and granular visibility we can prevent degradations, automate actions for improvements, and better communicate with our customers." Mahesh Anjan, Senior Product Technology Executive. AT&T. Watch webinar on demand.
The acknowledgements section is your opportunity to thank those who have helped and supported you personally and professionally during your thesis or dissertation process. Thesis or dissertation acknowledgements appear between your title page and abstract and should be no longer than one page. In your acknowledgements, it's okay to use a more ...
In preparation for that event, CAOE is soliciting the research community for white papers covering innovative supply chain resilience research ideas and solution methods as they relate to Homeland Security. ... White Paper Requirements and Acknowledgements: White Papers are limited to at most two (2) pages in at least 11 pt font; ...
Parkinson's disease is lacking easily accessible biomarkers. Here the authors show, that targeted blood proteomics is feasible to identify the patients and to predict the phenoconvertion in ...
The loss of electronic medical records has seriously affected the practical application of biomedical data. Therefore, it is a meaningful research effort to effectively fill these lost data. Currently, state-of-the-art methods focus on using Generative Adversarial Networks (GANs) to fill the missing values of electronic medical records, achieving breakthrough progress. However, when facing ...
Preferred orientation of macromolecules is one of the major issues that are commonly encountered in obtaining isotropic cryo-EM maps. Here, a comprehensive examination was performed of how macromolecule orientations respond to changes in physical factors, such as freezing temperature, and chemical factors, such as the addition of surfactants, for a standard set of macromolecules, which ...